To create an individualized cancer vaccine, researchers must identify cancer-specific peptides called neoantigens, then use a cell-, protein-, or nucleic acid–based platform to deliver those neoantigens to patients to prime the immune system to attack the tumor.
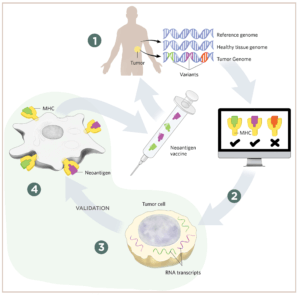
DNA sequencing, alignment, and variant calling
Next-generation sequencing data from tumor and normal DNA are aligned and compared to the human reference genome and then to each other to identify tumor-specific alterations. These variants are then evaluated for their resultant changes to the amino acid sequences of the encoded proteins.
Epitope prediction
The selected sequences are evaluated by computer models that predict the binding of the neoantigens to the major histocompatibility complex (MHC) proteins that would present them on the surface of cells.
Candidate Neoantigens Filter #1
RNAseq data from tumor RNA are evaluated to ensure the predicted alterations are being made into RNA transcripts, and we can further cull the list for other reasons, such as lack of sequence coverage of that region or gene.
Candidate Neoantigens Filter #2
Researchers isolate MHC proteins from patients and evaluate their bound peptides by mass spectrometry to validate that these peptides are being presented by MHC. In aggregate, such data can be used to improve the computer models that predict neoantigens.
Read full, original post: Infographic: Neoantigen Prediction for Personalized Vaccine Design