Gene Editing / CRISPR
Gene editing, or genome editing, is a new type of genetic engineering that is revolutionizing human medicine and agriculture. These techniques give scientists the ability to more precisely modify the genome of almost any organism. Scientists can pinpoint and remove genes that cause rare diseases in humans or insert desired traits like drought and disease-resistance already found elsewhere in a plant species [For more, read the GLP’s FAQ on gene editing in agriculture here].
Although gene editing can involve transgenics—the moving of genes from one species to another—it usually does not, diminishing the criticism from some anti-biotechnology experts who believe transgenics violate the ‘natural order’.
Some types of gene editing, such as the popular CRISPR-Cas9 system, have been likened to a “biological word processing system” that allows scientists to cut and paste DNA almost as easily as if they were editing a journal article. The gene editing revolution has just begun, and already it’s led to innovations. Many scientists believe gene editing has the potential to solve some of the world’s most pressing disease and agricultural challenges. Others worry that this new technology could be co-opted for malevolent purposes or that unintended consequences could arise.
The CRISPR system has garnered much attention for its ability to more accurately and efficiently insert and turn off desired traits, curing genetic diseases and increasing crop yields. It is also heralded as the cheapest, quickest and easiest to use form of genetic engineering to date. CRISPR, which stands for Clustered Regularly Interspaced Short Palindromic Repeats, is a natural bacterial defense system. In popular usage, “CRISPR” (pronounced “crisper”) is shorthand for “CRISPR-Cas9.” Found in a range of bacteria, CRISPR defends cells by identifying the DNA of invading viruses and, together with a protein made by bacteria (Cas9), slicing parts out of the virus to deactivate it—like a pair of DNA-cutting scissors.
Scientists have successfully “programmed” CRISPR to target and edit DNA at precise locations. The technique can be used to alter the genome of almost any organism. CRISPR allows scientists to delete and replace genes. When the DNA strand is broken, the cell’s repair systems kicks in to repair the break. Scientists can take advantage of this process by inserting new sequences of DNA at the repair site, thereby changing the gene sequence.
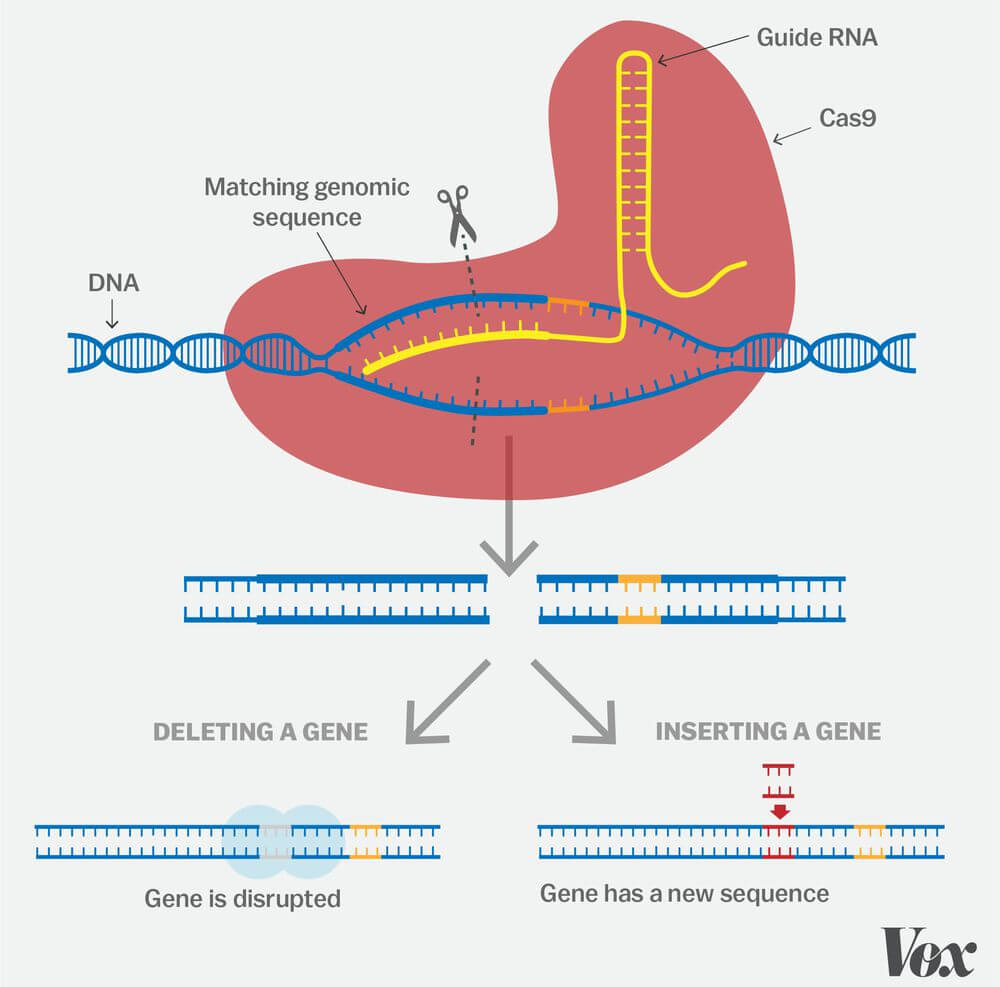
Credit: Vox
In the video below, Feng Zhang, a neuroscientist at MIT who played a central role in the development of CRISPR, explains how it works:
#Crispr can be applied in different ways w/ different outcomes which require differentiated regulations. Plants should not be subject to biotech regulations if they could also be obtained through earlier breeding methods or result from nature. @Food_EU @eaAgriFood @POLITICOEurope pic.twitter.com/Wd1PwVZfYH
— Euroseeds (@EuroseedsEU) July 25, 2019
Recent breakthroughs include an entirely new CRISPR-based gene editing tool that targets RNA, allowing for transient changes to genetic material, and a more refined type of CRISPR gene editing that can alter a single DNA base letter. Other CRISPR systems, such as CRISPR-Cas1, are also being developed.
Other popular forms of gene editing include zinc finger nuclease (ZFN), transcription activator-like effector-based nucleases (TALEN), and meganucleases. ZFNs are artificial restriction enzymes that can be engineered to target specific DNA sequences. TALENs are restriction enzymes that can be engineered to cut specific sequences of DNA. Meganucleases are homing endonucleases that can be used to replace, eliminate, or modify target sequences of DNA. Investigation of these systems predates CRISPR by years, but CRISPR has grabbed the spotlight because it is so much more efficient.
With gene and genome editing systems, researchers can permanently modify genes in living cells and organisms, including in human embryos, and, in the future, may make it possible to correct mutations at precise locations in the human genome in order to treat genetic causes of disease. Studies using laboratory and animal models of human disease have demonstrated that the technology can be effective in correcting genetic defects linked to single gene mutations that are correctable. Examples include cystic fibrosis, cataracts and Fanconi anemia.
Around 20 trials in humans have already begun or will start soon. Almost all of these remove cells from an individual’s body, editing the DNA and then putting the cells back into the body. In August 2017, researchers used CRISPR to correct a disease-causing mutation in dozens of viable human embryos for the first time. The researchers targeted a mutation in a gene called MYBPC3, which thickens the heart muscle—a type of hypertrophic cardiomyopathy that can cause sudden death in young people, especially athletes. The experimental procedure—the embryos were never implanted into a woman’s uterus—controversially changed the human germline. Many gene editing skeptics on the religious right and the activist left are openly critical of making changes that impact future generations.
In November 2017, the first human was injected with a gene-editing tool. Scientists infused the person’s blood with ZFNs, aiming to treat a rare metabolic disorder called Hunter syndrome.
CRISPR has also been used in the quest to transplant livers, hearts and other organs from pigs into humans. Researchers used CRISPR to create gene-edited piglets cleansed of viruses that might cause disease in humans.
[For more, read the GLP’s FAQ on gene therapy here].
Plant geneticists have used CRISPR to create a variety of novel changes to crops and farm animals, including virus-resistant pigs, disease-resistant cassava, low-gluten wheat (that people with celiac disease could eat), non-browning mushrooms, low-fat pigs that better regulate heat (which would better protect piglets from cold weather, a common cause of death) and oilseed crops with high levels of omega-3 fatty acids. Scientists have used another gene editing technique, TALENs, to create hornless cows (which make it less likely that cows harm each other and their handlers), soybeans with higher quality oil and disease-resistant rice.
Gene editing is similar to conventional breeding, but faster, cheaper and more precise. One of gene editing’s main advantages in agriculture is that it may be less strictly regulated than “first generation” genetic engineering, commonly referred to as GMOs.
This is because gene editing is more accurate, causing fewer “off-target effects,” and because it allows researchers to move genes from within the species, while GMOs often rely on genes from other species.
A gene drive is a natural phenomenon whereby the inheritance of a particular gene or set of genes is favorably biased, resulting increasing its frequency in the population. Gene drives can arise through a variety of mechanisms, and scientists have proposed using gene editing to engineer gene drives for specific purposes. These include preventing the spread of insects that carry pathogens, such as mosquitoes that transmit malaria, dengue, Zika and other diseases.
Here is how it works: This system uses genetically modified male mosquitos to deliver new genes along with a mechanism for copying the new sequences from one member of a chromosome pair to the other. In other words, a mosquito larva has a gene that came only from its father, yet has it as both a paternal and maternal copy. Thus, even a recessive gene will manifest its trait in all offspring. Furthermore, the offspring will spread the gene and trait to their own offspring. Since mosquitoes have a short life cycle, this means that in the course of just a summer, we could alter almost the entire population of a particular mosquito species in say the Brazilian rain forest, possibly wiping out the Zika disease. In August 2016 the U.S. Food And Drug Administration (FDA) issued a “Finding of No Significant Impact” to biotech company Oxitec’s plan to release genetically modified male Aedes aegypti mosquitoes into the Florida Keys.
Engineered gene drives have also been proposed to control invasive species, such as rodents that eat the eggs of endangered bird species in New Zealand, and for eliminating herbicide and pesticide resistance in crops.
Concerns about gene drives include the possibility that a mutation could happen during the engineered gene drive, which could spread unwanted traits with the drive. The spread of some other disease could be unexpectedly facilitated. Or the elimination of a link in the food chain could harm the local ecology. It’s also plausible that something could happen akin to the introduction of rabbits in 19th century Australia, where the population exploded, due to lack of predators, with major consequences for the ecosystem. There are also worries that an engineered gene drive could move beyond its target population, causing unintended impacts on other species and ecosystems.
Anti-biotechnology activists including Vandana Shiva, Jane Goodall and David Suzuki have advocated against the use of gene drives. In August 2017, they joined with other radical environmental groups to issue a well-publicized opposition statement to gene drive technology, writing:
Given the obvious dangers of irretrievably releasing genocidal genes into the natural world, and the moral implications of taking such action, we call for a halt to all proposals for the use of gene drive technologies, but especially in conservation.
In 2016, the National Academy of Sciences issued its wide-ranging review of dozens of studies, Report on Gene Drives in Non-Human Organisms, which outlined a number of potential risks but urged more research and gave a cautious green light to “highly controlled field trials.” Some studies have come to different conclusions, among them: researchers at the University of California, San Diego and colleagues at Harvard created a mathematical model for CRISPRs likely success, concluding the a gene drive could be remarkably aggressive in the wild, spreading a new gene beyond its targeted population—possibly meaning that experiments in the real world are too risky on a case by case basis at this stage in the technology’s development.
Do-it-yourself biology, also called “biohacking” or “DIY bio,” is a movement in which people are experimenting with biotechnology research and development methods outside of traditional research institutions. Some “biohackers” are trying to make these methods easier and more accessible, so that even non-scientists can use them. Because of its relative ease to deploy, CRISPR experiments can be performed even by high school students.
One central aspect of the DIY biology movement has been the sharing of laboratory space and equipment, as well as data and methods. One of the most accessible forms of biohacking is through engineering microorganisms or plants. Experiments range from using plasmids to create fluorescent bacteria, controlling gene expression using light in bacteria and even using CRISPR to engineer the genomes of bacteria or yeast. Some biohackers have begun selling kits that allow you to use CRISPR at home. One kit, created as part of an Indiegogo crowd-funding project, was sold for $130 by biohacker Josiah Zayner.
There are concerns that CRISPR might inadvertently alter regions of the genome other than the intended ones. These are called “off-target effects.” The worry is that CRISPR could change a beneficial gene, such as disabling a tumor-suppressing gene or activating one that causes cancer. Another concern is that because no two people’s genomes are identical, identifying off-target effects in individuals may be impossible. Researchers attempt to predict where in the genome off-target effects might occur using web-based algorithms, but there are concerns that this approach is not accurate enough.
In May 2017, an article published in the journal Nature Methods reported an alarming number of off-target mutations in mice whose genomes had been edited using CRISPR. However, experts voiced skepticism of the finding because only two mice were edited and unusual methods used. Scientists are attempting to address these concerns by developing more precise variants of the Cas9 enzyme used in the CRISPR system. Some of these enzymes have been shown to improve targeting in human tissue in the lab. Researchers have also focused on developing methods to more efficiently locate off-target mutations in the animals they study.
Chinese cloning company: Human clones probably already possible

Embryonic stem cells can help reverse blindness
Sharp restrictions on human gene editing counterproductive

James Watson: Basing ‘war on cancer’ on genome research diverts resources
Using CRISPR to edit crops ensures no transgenes, undercuts key anti-GMO ‘foreign gene’ criticism

International CRISPR summit ends in human genome editing slow down
What is actually possible with gene editing technologies?
How CRISPR works, according to its discoverers
Does germline gene editing cross evolutionary lines?
Can CRISPR be made even more precise?
Gene edited mosquitoes could wipe out malaria forever

Gene editing opens possibilities for improved livestock, but regulation controversies remain
Animal biotechnology regulation needs overhaul

CRISPR/Cas9 gene editing improved meat volume and cashmere production in goats
Will gene drive safety concerns limit breakthrough research?
Which of our 25,000 genes matter most?
Geneticists aim to eradicate malaria with genetically engineered mosquitoes
What do experts have to say on dangers, merits of gene editing?
Language matters when reporting gene editing stories
Biohacking pioneer Ryan Bethacourt on why genetic engineering is tool for progress
Biology ‘designers’ build new organisms by DNA swapping
Can insects be genetically engineered as bioweapons?

Regulatory breakthrough in Europe? Sweden says some gene edited plants not GMOs
Bio-safety measures can keep gene drives from causing ecological havoc

CRISPR Cas9 genome home editing kit brings genetic engineering to hobbyists
Gene editing safety regulation threatened by increasing ease of technology


