Type the words ‘e coli’ and ‘gmo’ into Google and you get a number of worrisome hits. Could your GMO corn be giving you a dangerous illness? When you drink diet sodas are you drinking bacterial poop? Activists group have frequently played up the alleged dangers of what they call E. Coli “contamination.
Activist groups have frequently raised fears of what they call E. Coli“contamination.
“E-coli live in the gut, and are beneficial,” writes Barbara Peterson, who is neither a farmer nor a scientist, on the site Farm Wars. “That is, unless we ingest one that has mutated into a harmful strain, and then it can cause disease.” Petersen goes on to write:
A little-known fact is that during the genetic modification process, e-coli is used to clone the transgenic (genetically modified) DNA before it is inserted into plant cells that are then grown into transgenic crops such as those bearing the label “Roundup Ready.” It is possible that a mutated form of e-coli resulting from the cloning process used in creating GMOs could get into the gut of a person or animal that eats a transgenic plant. Could this be why DuPont, the company that produces Pioneer Hi-Bred GMO animal feed, and proud proponent of GMO technology, has jumped into partnership with the USDA in identifying hard-to-identify strains of e-coli? With this convenient partnership, even if harmful strains of e-coli relating to GMOs are discovered, it is likely that the public will never hear it from the USDA or DuPont.
In the immortal words of George Costanza from Seinfeld upon swallowing a fly — “What can happen?!” The notorious bacterium Escherichia Coli is indeed used extensively in biotech; but is it a danger to people who might consume GMO foods? I wanted to know more about how and why E. Coli ended up becoming the bacterial lab rat. Let me tell you what I found.
Guts
In the late 1800’s scientists had begun exploring a largely unknown habitat. Even though it seemed inhospitable they found that it was teeming with life. One man, Theodor Escherich, discerned minute rod-shaped creatures swishing around there; based on their similar shapes and movement, they were given a single name after their discoverer — Escherichia Coli. The habitat of course was our own guts, and these rod-shaped creatures, which turned out to be a type of bacteria, were first detected in the feces of healthy breastfed infants.
Now we know more about them, and it turns out that the grouping known as ‘Escherichia Coli’ is quite a disparate one even though they look alike. No more than 20 percent of genes are shared by all, and two of them might only share 75% of their genes. For comparison, humans share about 85% of their coding DNA with the mouse. The E. Coli we house in our guts are beneficial ones that produce vitamin K for us (they don’t do this for free — we give them food and shelter). Some others, on the other hand, like the strain that goes by the name O157:H7, can cause dysentery. Even though this phrasing is extremely common in the media, saying that someone contracted a case of E. Coli is about as precise as saying that Jane Goodall once lived among mammals.
The comparison with mammals is an interesting and deep one. Our lower guts are the E. Coli’s main habitat; and when I say ‘our’, I mean us mammals. Our common ancestor with all other mammals probably lived around a hundred million years ago and looked a lot like a rat or shrew. As we have diverged from that ancient shrew into various shapes and sizes and spread all over the world, so have our E. Coli fellow travelers. As we get used to different diets and environments, the E. Coli in our guts evolve to thrive in the food bath that comes down to them. The host’s particular temperature, acidity level, salinity, diet — and even whether the animal is an urban or wild one — matters to their E. Coli hitch-hikers. The E. Coli genome has evolved alongside the mammalian one.
E. Coli outbreaks
In May this year 16 people got sick, some hospitalized, from an outbreak of E. Coli illness that was ultimately traced to a batch of clover sprouts. Another outbreak at the same time was related to ground beef and made 12 people sick. Last year, 33 people were made sick with ready-to-eat salads. What connects these outbreaks? Why salads sometimes, ground beef other times?
What connects them is the Shiga toxin. It acts in ways similar to the ricin poison and causes gastrointestinal sickness in humans. There are only a few strains of E. Coli that can produce cShiga — the most prominent one is O157:H7, a normal resident of cattle guts. Given that fact, and given that cooking kills it, you can probably imagine the scenarios that might cause an outbreak. One scientist in the papers I read helpfully mentioned that humans don’t usually get this infection directly from cattle gut (‘directly?’) But undercooked beef might certainly harbor it; so could a manure application to growing vegetables that has been incompletely composted. E. Coli from infected manure can be drawn into leaf veins, so simple washing may not get rid of the contaminant.
The Shiga toxin is a well-constructed cell-killing machine made of sub-units that each help in different parts of the process. One sub-unit helps it attach to its victim cell and penetrate it. Once it has invaded a healthy cell, another sub-unit is released that cripples RNA function, causing cell death. Repeated over many cells, for us this means diarrhea; dysentery; possibly kidney failure depending on how far the Shiga-toxin producing E. Coli have reached.
Lab rat
So that’s the bad guy, now let’s talk about some good guys. One that goes by the name of E. Coli K-12 has lived in our labs for almost a hundred years. It may not be particularly pleasant to think about but K-12 originally came from the stools of a diptheria patient in Palo Alto. Since then, it has become a pet of the microbiologists and better understood, down to its genes, than almost any other creature. Over time, it has been bred, cloned, infected with phages, cured of phages, bombarded with X-rays, UV rays, doused with chemicals, and basically been turned into a friendly dog from the wolf that it used to be. What I mean by that is that K-12 has been domesticated; it can no longer live in its wild home, the human gut. The lab is its new human gut.

At this point, K-12 is given the Biosafety level of 1 — the safest. From the Wikipedia definition, this means that it is “not known to consistently cause disease in healthy adult humans, and of minimal potential hazard to laboratory personnel and the environment (CDC,1997)”. Some oversmart scientists have demonstrated this by sipping E. Coli K-12 cultures. This is never a good idea, but what gave them the confidence to do so was the knowledge that K-12 cannot survive outside the lab for long — it would soon be dead in their guts. Also, it never did produce Shiga or other toxins since it lacks the mechanisms to do so.
Following grand E. Coli tradition, the relationship between K-12 and the microbiologists is a symbiotic one. We spend many research dollars on it to make sure it is safe and reproduces (when we want it to) and lovingly feed it agar-agar. While K-12 for its part has been instrumental in our understanding of how DNA works, and many other fields of study.
Cut and paste
We all use computers, but how many of you hack them? I use Microsoft Windows every single day and it often does one or other annoying thing. But I have neither the ability, nor the tools or expertise needed to go in and ‘fix the code’.
We were in pretty much this position with living cells about half a century ago. Scientists knew that each living cell contained a double-helix — its DNA. And that the sequence of letters in it programmed the cell’s functioning. But they did not and could not manipulate those gene sequences. From that to the DNA hacking world we live in today: how did we get here so quickly? Pardon the awkward metaphor, but we rode in on the shoulders of E. Coli K-12.
In fact the very first such DNA manipulation ever done involved taking some DNA from one E. Coli and putting into another, using a bacteria-infecting virus almost like a pair of pincers. Infect one bacterium, pick up some DNA into the virus, infect another bacterium with the same virus, pass the DNA on. This became known as recombinant DNA.
The period from the late 60s to the seventies saw many separate teams working on techniques to create ‘chimeric’ DNA — those that contained sequences from different species. Some researched restriction enzymes — these are enzymes that can cut DNA into bits at precise points, almost like a pair of scissors that will only cut at previously marked lines. The points are defined by their gene sequences. Some researched DNA ligase — a sort of glue that nature uses to repair broken DNA.
Cut, then paste. In some ways chimeric DNA is like a collage, and one must do things that one does to make a collage. Cut, then paste, and then — of course, make many copies to distribute. This is where E. Coli K-12 came in. It was, and is, used in biotech as a copy machine.
Copy Machine
The biotech industry officially kicked off with a patent awarded to Stanley Norman Cohen and Herbert Boyer for their technique of recombinant DNA (ominously, it seems that the patent was quite a bit broader than it needed to be).
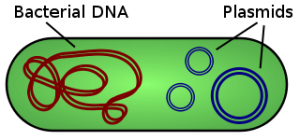
Stanley Norman Cohen was researching plasmids. Plasmids are circles of DNA that exist independently of the main DNA of the bacterial cell. If the main DNA is iOS that runs your phone, the plasmid might be a separate app that you purchase for a specific purpose. Being DNA, it can replicate, and it does replicate, inside the cell, at a rate that can be faster than the rate of the bacterial replication itself. One of the functions of plasmids is to carry gene sequences that give antibiotic resistance to its host for one or other antibiotic.
Meanwhile Herbert Boyer‘s lab was researching a restriction enzyme that would not only cut the DNA in specific places, but also leave sticky ends, making it easier to join up later.

The very first genetically engineered product that came out of this was human insulin. It replaced the use of insulin from cows or pigs, which caused immune reactions in some. Let’s see how the E. Coli copy machine worked. The insulin gene was isolated from humans. Then came the cutting and pasting techniques described above.
- An E. Coli plasmid is first cut using a restriction enzyme — now it has open ends and is no longer a circle.
- The insulin gene is attached to one end using the paste technique with ligase.
- Then the circle is joined back up with ligase. This time it contains cargo — the human insulin gene.
- The plasmid is inserted back into the bacterial cell using heat shock — a technique developed in 1970.
- The fact that antibiotic resistance is also given by the plasmid makes it easy to recognize the cut-and-paste plasmids from the untouched ones. If the insulin gene is joined at the hip with the gene for antibiotic resistance, and if the bacterial colony that has been grown is treated with the

Eli Lilly & Co. insulin manufacturing site antibiotic — those that are killed off are the ones that didn’t have that antibiotic resistance gene. Those that survive carry the resistance gene and also its partner the insulin gene.
- The E. Coli that are surviving are the hijacked ones, but they themselves are none the wiser. We feed the E. Coli; they multiply; out come a lot of E. Coli, and with them, out come many copies of the human insulin gene.
- What genes like to do, is make protein. This they do, as a matter of course. Therefore, the hijacked E. Coli make human insulin.
GM plants
Now if it weren’t the insulin gene, but rather some gene that needed to go into a plant — let’s say, to take a wild (but perhaps not implausible) example, a gene that makes flowers glow in the dark — the gene would have still further to go. We start with the gene being cloned many times in E. Coli as above, and continue from there:
- We now have many copies of the glow-in-the-dark gene ensconced in a plasmid.
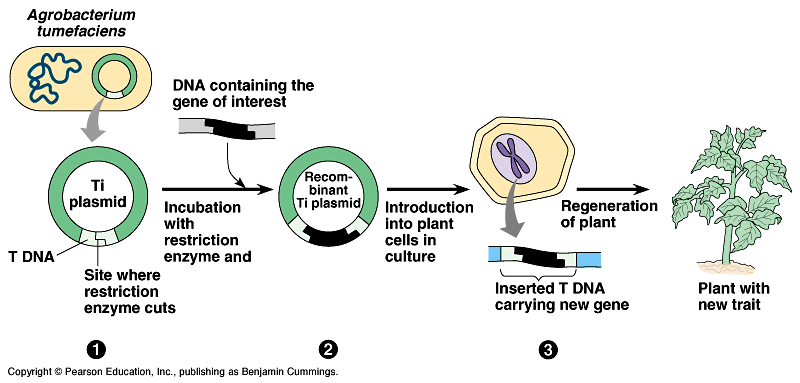
- This is inserted into yet another bacteria using the same cut-and-paste plasmid technology. This time, it is the Agrobacterium tumefasciens — bacteria that has the special skill of being able to insert its plasmid into plant cells.
- The plasmid with the glow-in-the-dark gene goes into plant cells
- The plant is grown from a culture of cells
- It makes seeds. These seeds contain the desired gene and will grow plants that exhibit the glow-in-the-dark behavior.
- By this point, we are quite distant from the use of E. Coli.
School of Biological Sciences, University of Texas
Safety and the lack thereof
Regardless, do you still worry about the what-if’s and even-so’s of using microbes in the lab? You will be happy to know that you are not alone. Something rather astounding happened in those early days of recombinant DNA, perhaps even more astounding than the experiments with life that were ongoing.
That was this. People raised the alarm about the sheer power of the experiments they were performing in the lab and willingly stopped all work on DNA entirely until an elite community could get together and think a little. The astounding thing was that this wasn’t activists raising the alarm, nor journalists, or governments, or doctors — but scientists themselves; and not rival scientists throwing sand in the wheels to stop to experiments they didn’t want to see succeed, but rather, the very scientist who was performing the work.
This was Paul Berg, one of the originators of recombinant DNA technology, who later went on to win a Nobel prize. He organized a conference in Asilomar, California in 1975 to take stock of potential dangers of biotechnology. Meanwhile all concerned scientists observed a complete moratorium on such work. What came out of it were a set of safety guidelines from NIH that have been improved over the years and are still followed in biotech labs. We are long past the wild west days of biotech.
More importantly, the conference made their work transparent and included the media in their discussions, thereby ensuring that decisions weren’t made behind closed doors. Sounds like a great model for today.
Given all the above, the use of E. Coli in the lab seems very very safe to me, and no one has ever known to have gotten sick from, say, recombinant insulin. But ultimately the best way to get a grasp on safety might be in a relative way.
We routinely accept the much greater risk of using manure in farming. This manure is from the cattle gut, which is where the harmful strains of E. Coli actually do live. Plants can take up E. Coli of the harmful sort and unless they are cooked, can cause illness. There are, once again, regulations about having to completely compost the manure before use, and time limits on how long before harvest it can be applied. Regardless, E. Coli sickness from bad farming practices have known to have happened, though they are rare. In contrast, no one has ever gotten E. Coli sickness from GMO foods or insulin.
Aneela Mirchandani is a software engineer by profession, a writer and biology geek. She blogs about food at The Odd Pantry. Her work has appeared in Harpur Palate and The Indian P.E.N. You can follow @theoddpantry on Twitter or email her at [email protected]
Additional Resources:
- Personal Reflections on the Origins and Emergence of Recombinant DNA Technology, Genetics
- E.coli’s use in GMOs: can you get E.coli poisoning? Biofortified
- E. coli – the biotech bacterium, Biotechnology Learning Hub




