Seinfeld episode entitled “The Betrayal” original air date November 20th, 1997:
Usha Ranawat: Don’t go! India is a dreadful, dreadful place.
Zubin Ranawat: You know it’s the only country that still has the plague. (laughing) I mean the plague! Please!
Someone should tell the Ranawats that if the definition of a “dreadful, dreadful place” is the presence of the plague, that means the United States is a pretty dreadful place too.
Many people will read The Plague, or The Black Death or The Bubonic Plague and and conjure up images of plague doctors, the Middle Ages and “bring out your dead”. It is most notably seen as a disease of the past, and most certainly it is one that is over. We all are guilty of mislabeling this diseases as simply a 14th century pandemic. While this is true, the black death’s causative agent, the bacteria Yersinia Pestis, never really left us–and it’s alive and well here in the United States.
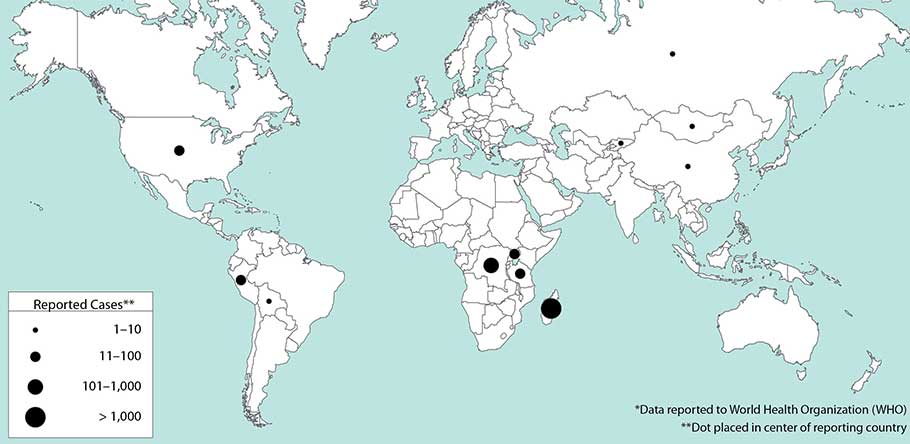
Over the last few decades an average of seven cases of the plague a year have occurred in the U.S., and coincidentally since 2000, India and the United States have had about the same number of cases. Most of the cases here occur in the western United States, particularly in the southwestern states of New Mexico and Arizona. Already this summer two people have died of the plague in Colorado and a child has contracted the infection at Yosemite National Park where rangers are working tirelessly to control an outbreak among wildlife.
But our relationship with Y. pestis is not just about a rising body count, like any two organisms that interact we have influenced each other’s genome and each other’s evolution in significant ways.
The Plague’s toll
It not should be surprising that organisms affect each other’s genome and therefore their evolution. We have evolved some resistance to the malaria parasite most notable through the increase in the frequency of the sickle cell trait gene; the same can be said of the tuberculosis bacterium with the cystic fibrosis gene. It therefore stands to reason that the Y. pestis has had some affect on our genome, however, the extent of which has been elusive until recently.
Romania has provided an excellent backdrop to study the impact of the plague on our genome. Culturally, it is made up of two distinct ethnic groups: European Romanians and the Rroma (more commonly known as gypsies); the former has been in the region for approximately 2,000 years, while the latter arrived about a millennia ago from northern India. Much like the rest of Europe, Romania was marred with frequent plague outbreaks during the 14th century. The same cannot be said of India during this period.
By comparing the genomes of modern day Rroma, Romanians, and northern Indians researchers have been able to investigate our evolution from plague exposure. Any changes that are shared by the Rroma and Romanians (both of which were exposed to the bacterium) but are absent in the Indian, (where it did not hit) are potential targets. Contemporary comparisons of the genome have found that about 20 genes show distinct similarities in the Rroma and Romanians that are absent in the genome of northern Indians. Of these, several appear to be inconsequential to plague exposure like genes for skin pigment and a gene implicated in a neurodegenerative disorder.
However, a set of three genes on chromosome 4 that code for proteins associated with the innate immune response proteins called toll-like-receptors (TLRs) have piqued researchers interest. TLRs act just as their name implies: as a “toll”, they sit on the surface of many immune cells and are activated by broad classes of antigens (e.g. the lipopolysaccharide of gram negative bacteria). When activated, the cells alert cells B and T cells which can launch more specific responses to the pathogen.
The Rroma and Romanians shared the same three TLR sequences (i.e. alleles), which varies greatly from the 3 TLR alleles of the people from northern Indian. Furthermore, the Rroma and Romanian alleles are very similar to those in other people with European dissent. However, amongst those with Asian and African dissent, where the plague was not endemic, these plague specific TLR alleles are no where to be found. Cell culture tests have shown that varying the sequences of these genes provides different intensities of immune response to the presence of Y. pestis. So it appears that these three TLR sequences increased in frequency in the European gene pool due to their ability to quickly, effectively, and vigorously alert the immune system to the presence of the plague than other sequences.
Emergence of a pest(is)
In evolutionary history,Y. pestis is a relatively young pathogen. Genetic evidence suggests that it evolved from Yersinia enterocolitica (also known as Yersinia pseudotuberculosis) which causes yersiniosis, a mild gastrointestinal infection, at some point in the last 10,000 years. In this short time,Y. pestis has acquired several genetic changes both chromosomal and extrachromosomal that have morphed it from a mild self-limiting disease to one of pandemic proportions.
By starting with the ancestor Y. enterocolitica, scientists have been able to either add or subtract genes to elucidate what changes were necessary for speciation to occur. Interestingly one of the common themes in the its evolutionary history is that Y. pestis has frequently lost genes with the result being increased virulence. Lipopolysaccharide (LPS) is one example of this. The LPS, also referred to as an endotoxin, is one of the most significant virulence factors in gram negative bacteria (a group that contains Salmonella, Shigella and E.coli). LPS frequently causes septic shock, an often fatal condition in which the immune system overreacts to the molecule’s presence.
A major part of the LPS is the o-antigen (you may recognize this from the food-borne pathogen E. coli “O”157). As deadly as this molecule is, the plague bacteria doesn’t express it. The genes that synthesize the o-antigen are present in both Y. pestis and Y. enterocolitica, however, of the 17 genes involved in synthesizing the o-antigen, five are disrupted (four via deletion; one via insertion) in Y. pestis which inhibits the gene’s expression.
For scientists, the hardest evolutionary question to answers has been: how did this bug that as Y. enterocolitica was spread via fecal-oral route evolve into one that spread through both the infamous “fleas on rats” mechanism and respiratory secretions. Scientists have now pinned down that just a few minor changes to the genome were able to facilitate this change.Y. enterocolitica can survive in a flea’s hindgut, but has no transmissibility from there.
However, at some point in the bacteria’s evolutionary history it acquired a plasmid that allowed the bacteria to successfully colonize the fleas midgut. Then the loss of function in three genes allowed the bacteria to form biofilms inside the flea’s midgut. This ability was a key step in its ability to be transmitted through a flea bite, which again interestingly happened through the loss of genes. On the other hand to add transmissibility through respiratory secretions the plague bacteria picked up a plasmid that coded for just one advantageous protease.
In its short history with us, the plague bacterium has evolved significantly and has caused us to evolve as well. However, as with evolution in any species it is never done, both organisms will continue to shape each other.
Nicholas Staropoli is a research associate for the American Council on Science and Health. He has an M.A. in biology from DePaul University and a B.S. in biomedical sciences from Marist College. Follow him on twitter @NickfrmBoston









































