Advanced age and pre-existing conditions are clearly major risk factors for the development of severe COVID-19, however there are also large numbers of younger and generally healthy people who develop serious symptoms, as well as patients of all ages who rapidly recover, or never shows signs of the disease to begin with.
The number of viral particles an individual is exposed to is certainly one variable that may determine disease trajectory, as are environmental considerations, as well as access to treatment. Nevertheless, numerous studies are currently seeking to determine how a person’s unique genome may influence the ultimate impact of the disease. It is important to remember that in contrast to some bacterial and fungal pathogens that produce toxins which directly impact the host, most viruses like SARS-CoV2 do not, and the symptoms and ultimate cause of death are generally produced by the inflammatory responses of our own immune system.
Thus, studying how human genetics impact disease progression could both lead to a predictive understanding of who might require strict isolation, as well as who could be spared the worst of the disease impacts. In addition, if we know specific genetic variants that cause bad outcomes, this could lead to the development of selectively targeted drugs that treat at-risk individuals.

Identification of individuals with genetic resistance to infection, or who may be protected from developing symptoms, could help us understand and exploit vulnerabilities at the interface between virus and host, and may have epidemiological implications. Isolating asymptomatic spreaders could significantly reduce the prevalence of the virus in a population, while genetic variants that protect from initial infection could lead to development of preventative therapeutics that block viral entry into host cells, as well as provide peace of mind to the resistant individuals.
Putting COVID in genetic context
Many of the broader medical issues we face have some genetic basis. However, rather than resulting from single catastrophic failures in function, such as running over a nail and getting a flat tire, human disease generally arises from numerous subtle genetic and environmental variables. Mutations with a large negative impact are generally not present at high rates within a population.
Most human diseases with a genetic component are like driving off-road in an aging car with worn out shocks and bald, under-inflated tires that are out of alignment; something will eventually go wrong due to the multiple potential sources of failure placed in the context of particular environmental stresses. While debilitating genetic diseases caused by individual catastrophic mutations certainly exist, and can tell us a great deal about how body systems work, in the vast majority of human diseases, a combination of subtle genetic and environmental factors is to blame, not a single mutation in one gene.
Much of human disease, and thus most genetics research, is based upon the two alternative arms of this apparent dichotomy: rare mutations with major direct impact, or common variants with subtle relevance placed in particular environmental context. In the search for understanding how human genetics plays a role in COVID-19 disease progression, each of these paradigms is being investigated.
One potential limitation to the search for individual genes that cause significant impact on COVID-19 when mutated, is previous experience with a so-called “candidate gene” approach. This involves taking the knowledge of a disease and the genes that might be involved, and then searching for variants in those genes within individuals who display the symptoms of the disease. This is akin to rounding up the “usual suspects”, and often equally as unproductive.
The problem that can often arise with a candidate gene approach is one of causality. As our genomes are full of differences that have no detectable functional impact, if you look at enough people you are likely to find some variations that exist in one population when compared to another. This correlation may be a statistical “red herring” that doesn’t provide any real insight into the disease.
An extreme example illustrating the potential limitations of a candidate gene approach would be to compare two very genetically distinct populations that are subject to very different environmental variables. In such a case a variant identified in a candidate gene could simply have arisen randomly and may correlate with a difference in a disease incidence or severity, but in actuality has no direct functional significance. Science is self-corrective and subsequent studies attempting to prove causation can certainly be performed, however these can be costly and time consuming, and especially during an ongoing pandemic, this type of candidate gene approach might not make sense.
While investigating the variants of an individual candidate gene might not be the best use of resources, new advances in DNA sequencing technologies allow the low-cost and high-speed analysis of entire genomes in large groups of people. Whole genome sequencing was once cost prohibitive and took years of painstaking work to complete, but can now be performed in hours, for about the price of a nice laptop.
Furthermore, studies focusing only on the expressed portion of the genome, the so-called exome, can be undertaken even more efficiently, and can increase the odds that any mutations identified might actually directly alter function. Thus, genetic variants potentially relevant to disease progression can be identified across large diverse populations with greater statistical certainty.
COVID whole genome mapping ‘moon shot’
The COVID Human Genetic Effort includes a worldwide group of research centers coordinated by Dr. Helen Su from the National Institute of Allergy and Infectious Disease, and Dr. Jean-Laurent Casanova of The Rockefeller University and Howard Hughes Medical Institute. The overall goal of this project is to perform whole genome and exome sequencing from COVID patients who are not elderly and don’t have clear pre-existing conditions to look for mutations that might be responsible for good or bad outcomes. In particular, the researchers are searching for monogenic variants that either increase immunity and are protective, or lead to particularly severe disease in individuals that might otherwise be spared the worst impacts of COVID-19.
Importantly, a key aspect of this work will be to biochemically characterize the candidate variants to look for potential mechanism of action, and weed out genetic variants that might be randomly associated with particular disease endpoints. This project will employ cutting-edge sequencing and analytical techniques to look at large populations with a high degree of sensitivity across the genome, and include follow-up functional studies to directly identify causative mutations and avoid potentially faulty conclusions based solely on statistical correlations.
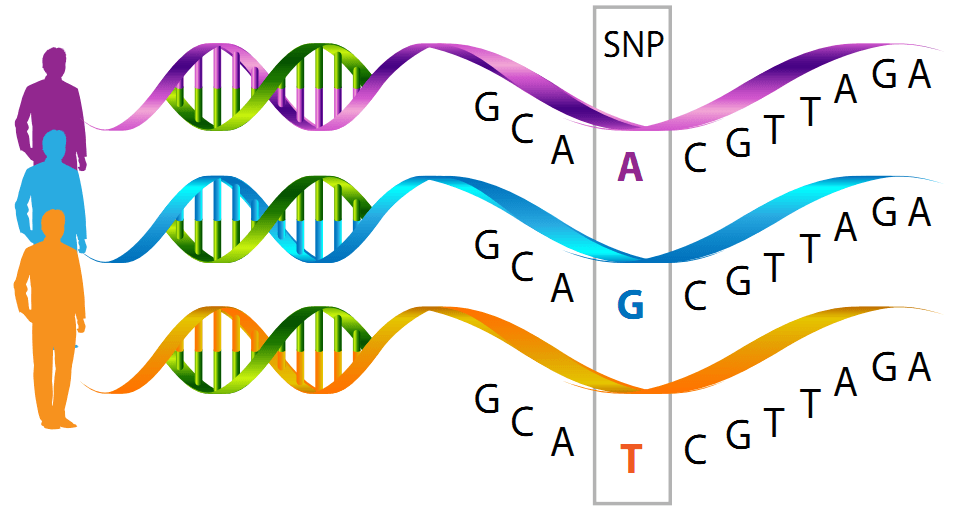
While identifying genetic mutations with specific functional significance can drastically impact our understanding of a disease and the potential search for beneficial therapeutic interventions, correlative studies of subtle genetic variations can also be extremely important. The term single nucleotide polymorphism (SNPs) refers to individual variations in genomic DNA that can be found within particular populations. The locations of thousands of SNPs in the human genome that can exist in one form or another are known, and rapid, powerful analytical techniques can easily provide a fingerprint of any individual’s personal complement of SNPs. Although individual SNPs don’t confer any obvious functional significance, there is tremendous power in being able to assess which SNPs a person carries across the genome.
SNPs are used to identify ancestry and personal genetic testing companies use panels which assess huge numbers of SNPs to determine one’s genetic background; for example, the part of the world from which a person’s family originated. Huge numbers of SNPs are looked at in concert across the whole genome and different patterns emerge which can then be correlated with the risk of developing particular diseases or pretty much any other characteristic with a genetic component. These analyses are broadly referred to as Genome-Wide Association Studies (GWAS).
The direct causative impact of any individual SNP is generally essentially zero, but the power is in the sheer magnitude of the numbers analyzed, both numbers of SNPs in one individual data set, and the combination across sometimes thousands of individuals. GWAS and similar types of analytical procedures can provide clues to the genetic basis of a particular disease process, and follow-up sequencing of genes harboring certain SNPs associated with a specific disease can provide further insight. Additionally, by treating a SNP signature associated with a particular condition as a “bio-marker”, predictive or diagnostic approaches can follow. In the context of COVID-19 this could mean identification of specific SNP patterns that might be associated with severity of disease or potential to respond to a particular therapeutic option.
Companies looking for genetic links to COVID susceptibility and recovery
The personal genetic testing company 23andMe, through the 23andMe COVID-19 Study, is currently conducting GWAS to look for associations between certain SNP patterns and COVID-19 infection, severity and recovery. PrecisionLife, a genetic analytics company based in Oxford, has announced the identification of 68 genes with SNP variants associated with severe COVID, including some that had been previously identified as playing a role in viral pathogenesis. PrecisionLife recently posted a pre-print on the website medRxiv which analyzed data obtained from the UK Biobank using an artificial intelligence genomic data analysis tool. The researchers stated that these results could have been obtained following traditional GWAS methods.
Whether through analysis of specific candidate genes, wholes genomes or exomes or GWAS with thousands of SNPs, a single study can only be so powerful, even if many individuals are included. Integrating numerous independent data sets into larger meta analyses can provide deeper and more meaningful understanding. This is where the COVID-19 Host Genetics Initiative hopes to make a significant impact.
Behind the scenes at the COVID-19 Host Genetics Initiative
Originally started by the Finnish Institute for Molecular Medicine, this multinational effort has been set up to share resources and data and organize multidisciplinary research activities. It includes clinicians, academic researchers and industry partners, such as DNA sequencing companies. One main aim is to create a series of agreed-upon phenotypic definitions and analytical procedures to harmonize data for meta analyses within an open science format in which results can be shared freely between partners. Although the potential for sharing data obtained with similar criteria and protocols holds tremendous potential power, international sharing of clinical data is very complicated and it will be interesting to see the results of this unprecedented effort—and extremely important in these unprecedented times.
The first results from these studies are starting to trickle in, and already there are examples of differential COVID-19 outcomes dependent on human genetics. People with blood group A seem more likely to develop severe COVID-19 with respiratory failure, while blood group O appears to confer a protective effect. A paper reporting these observations was recently published in the New England Journal of Medicine by the Severe Covid-19 GWAS Group. The study identified the ABO blood group locus and a cluster of genes on chromosome 3, including a region known to contain genes involved in immune cell function, associated with worse outcomes after analyzing ~1,600 patients in Spain and Italy with respiratory failure caused by COVID-19. However, another recent study in the journal Annals of Hematology found no association between ABO blood type and the severity of COVID-19 disease.

Hopefully, more results such as these will soon be published, and insights into how human genetics might impact COVID-19 severity will become evident. That could well lead to epidemiological actions and drive the development and deployment of therapeutic interventions. Until then, the more patients studied, and the deeper the analyses, the closer we will hopefully come to reigning in this terrible pandemic.
Joshua Z. Rappoport, Ph.D., is the executive director, Research Infrastructure at Boston College, and the author of the newly published Mapping Humanity: How Modern Genetics Is Changing Criminal Justice, Personalized Medicine, and Our Identities. Follow him on LinkedIn











































