Various innovations in the field of genomics over the past few decades have given researchers hope that resolutions to long-lasting debates might finally be on the horizon. In particular, many have become optimistic about the prospects for disentangling the threads of “nature” and “nurture” — that is, about determining the extent to which genes alone can explain differences within and between populations.
…
[O]ver the past two decades experts have come up with robust statistical techniques to address the issue, using data collected from thousands of individuals.…
[Two papers published in April] have illustrated that problems with interpretations of GWAS results are far more pervasive than anyone realized. The work has implications for how scientists think about the interactions between genetic and environmental effects.…
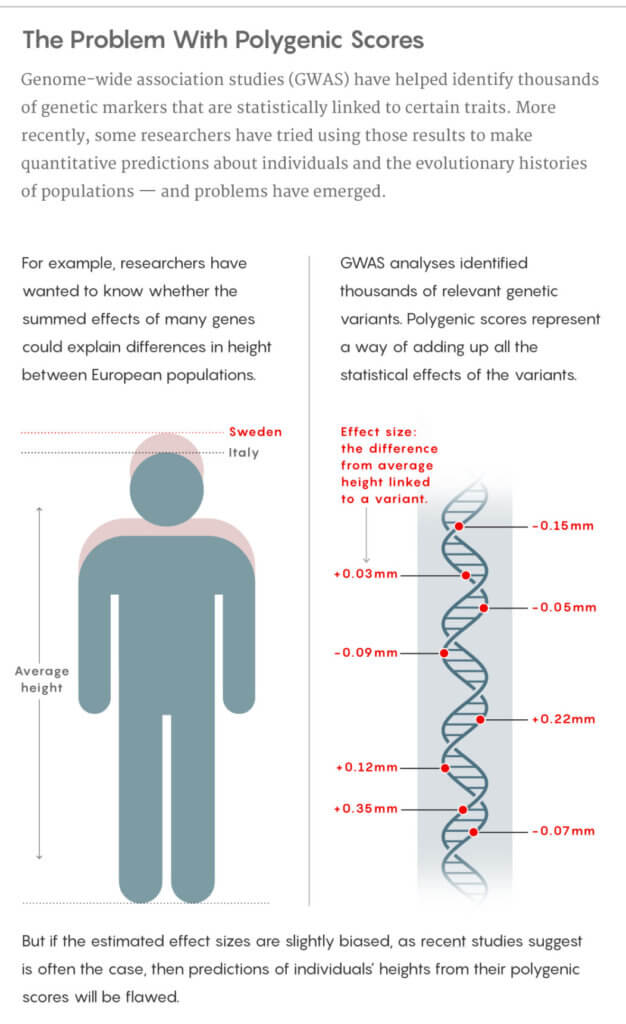
That’s not to say that genome-wide association studies do not have incredible power. They can still identify important markers for traits of interest. The small intrinsic biases don’t invalidate the thousands of regions of DNA such studies have already implicated in diseases and other conditions and traits. But the strengths of those individual and collective associations are general estimates. It’s when they’re accumulated to make inferences about differences between populations, both in evolutionary and medical contexts, that things can go wrong.
 Read full, original post: New Turmoil Over Predicting the Effects of Genes
Read full, original post: New Turmoil Over Predicting the Effects of Genes































