In the US, the Food and Drug Administration … GMO crops are deregulated once nutritional and compositional “substantial equivalence” is demonstrated. The set of parameters … necessary to declare a GMO as substantially equivalent … focuses on a restricted set of compositional variables, such as the amounts of protein, carbohydrate, vitamins and minerals. …
. . . .
Recent technologies used to ascertain the molecular compositional profile of a system, … collectively referred to as “omics technologies”, are used extensively in basic and applied science. … the majority of authors of [omics studies of crops] conclude that the statistically significant changes observed between the conventional and the GM varieties are not biologically significant because they fall into the range of variations … between different conventionally-bred varieties, and under different environmental conditions….
. . . .
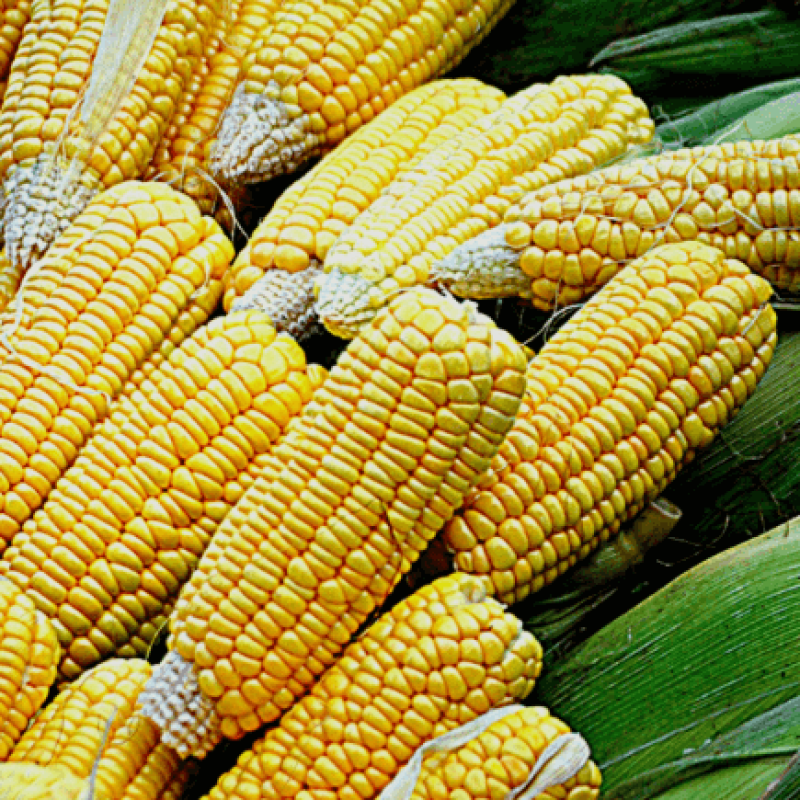
 …[W]e have performed proteomics and metabolomics analyses of NK603 (sprayed or unsprayed with Roundup) and isogenic maize kernels.
…[W]e have performed proteomics and metabolomics analyses of NK603 (sprayed or unsprayed with Roundup) and isogenic maize kernels.
. . . .
In this report we present the first multi-omics analysis of GM NK603 maize compared to a near isogenic non-GM counterpart. … Although NK603 had comparable nutritional and compositional profiles when originally accessed by the developer company upon registration of their product, our analysis … shows that NK603 grains, with or without Roundup spraying during cultivation, are not equivalent to isogenic non-transgenic control samples.
Several laboratory studies consisting of 90-day feeding trials in rodents have been conducted to evaluate the safety of GMO crop consumption. These investigations have frequently resulted in statistically significant differences in parameters reflective of disturbances in various organ systems and in particular liver and kidney biochemistry, but with interpretation of their biological significance, especially with respect to health implications, being controversial.
For more background on Seralini’s previous GMO studies and the funding for his research–this study was funded by an anti-GMO organic group, as listed in the paper–read the GLP’s Biotech Gallery profile. Editor’s Note: The GLP has asked independent scientists to review this study. We will update this comments as they come in:
Editor’s Note: The GLP has asked independent scientists to review this study. We will update this comments as they come in:
Seralini does not appear to understand the legal concept of “substantial equivalence”
1) There is no information as to whether he looked at compounds on the OECD list. These were chosen for their relevance to detect changes in key metabolic pathways. His list needs to be compared to the OECD one.
2) Once a difference has been found, the second step is missing altogether in this paper. Namely, all statistically significant, biologically relevant changes need to be compared to values in other varieties. If they are in the normal range, there is no issue. The actual differences observed are small, many much smaller than the differences found from one field to the next for the same crop. Statistically significant differences are not biologically significant differences.
3) At the end of the day, the OECD list covers about 60 metabolites that make up 95% of the grain. The remaining 5% contains many tens of thousand metabolites, where the “dose makes the poison” adage kicks in.
4) Differences in proteomes are for the most part irrelevant to food safety assessment. The reason for this is both simple and complex. Stated simply, large differences in proteomes are often seen in specimens with essentially identical compositions. The underlying complexity is that the concentrations of metabolite(s) produced by a protein or a set of proteins in a pathway reflect the interaction of a panopoly of variables such as enzyme concentration, substrate concentrations, concentrations of regulators, metabolic fluxes within the organism, and environmental regulation. Maybe a simple way to put it is that under the conditions at which a cell operates a great deal more regulates the concentration of a particular metabolite than the concentration of the enzyme that catalyzes its synthesis. I can’t tell you how many papers there are that report that enzyme X was elevated 2-fold or 5-fold or 100-fold but no change was observed in the product of its metabolic product. The rate limiting step in the production of a metabolite is not necessarily dependent on how much of the enzyme that produces it is present. Changes of a few fold are often inconsequential, and changes of a few percent sometimes evoke larger changes than one might expect. At the end of the day the proteome doesn’t tell us anything about safety. Proteomics can be a valuable research tool in the hands of an investigator with an hypothesis, but as applied here its simply another data dredging tool looking for a statistically significant change that means nothing.
5) What really counts from and safety and nutrition perspective is the metabolome. That’s the part we eat isn’t it? All of the changes in minor metabolites that were reported are small compared to the magnitude of changes in minor metabolites that is often observed in the same crop plant or its seeds. They are also meaningless changes; a few-fold change in polyamine concentration has no biological significance and if this experiment was done 10 times with NK603 the polyamine concentration would probably be all over the place.
6) There are some more fundamental problems with omics as an assessment tool which are covered in my 2010 paper. In particular, Lay et al published a compendium of troubles that face -omics as a science tool. These include the more obvious data dredging and statistics arguments to some pretty interesting math that indicates in some cases omics is more likely to give a wrong answer than a correct one. Think of it this way, the science of analytical chemistry has spent the last few hundred years developing methods for measuring one compound with accuracy, precision, reproducibility, sensitivity, etc. Omics is the emerging science of measuring everything poorly. When used correctly and in competent hands, omic analysis can be a powerful research tool. What it can’t do is tell you if two varieties of corn are equally safe to eat. It can, however, tell you that they differ in some way, and to an investigator in lab that may (or may not) be important to know, however, since there will no doubt be 100s if not 1000s of differences observed, good luck figuring out what they all mean. Omic analysis may eventually provide high quality useful data that tells us something about safety, but we are not there yet. Which of course means that the paper in question should be disregarded as artifact. Ir was in fact surprising to see that they did not observe more and larger differences.
The GLP aggregated and excerpted this blog/article to reflect the diversity of news, opinion and analysis. Read full, original post: An integrated multi-omics analysis of the NK603 Roundup-tolerant GM maize reveals metabolism disturbances caused by the transformation process