Both Neanderthals and Denisovans are distinct species in the genus Homo that coexisted and interbred with modern humans over 40,000 years ago. While some experts contend that Neanderthals and Denisovans fall under the same category as Homo sapiens, since together they satisfy one definition of “species” — a group of organisms whose members have the ability to produce fertile, viable offspring — the general consensus is that they are separate species.
Recent research suggests that human-Neanderthal hybrid males were not fertile, for example, while human-Neanderthal hybrid females were. In this way, our Neanderthal DNA may have been solely inherited from female hybrids.
All non-Africans today have a small percentage of Neanderthal DNA, while Denisovan DNA is particularly common in Asian and Oceanian populations (Australasia, Melanesia, Micronesia, and Polynesia). The idea of a “ghost population” — an unknown species inferred to exist through statistical techniques — had been postulated to account for archaic sections of the human genome that do not match Neanderthal or Denisovan DNA.
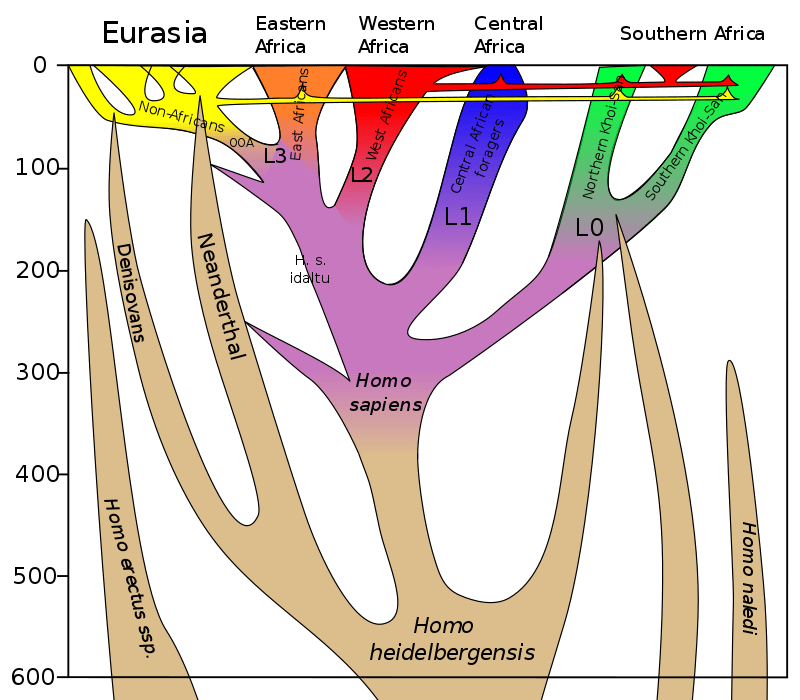
This third archaic species was identified using “a demographic model based on deep learning in an Approximate Bayesian Computation framework,” the researchers wrote in their article published in Nature Communications last month. “We have an overwhelming support for the existence of a third extinct branch of the Neanderthal-Denisovan clade,” they said.
The species appears to be responsible for a third unique introgression (the introduction of genes from one species to another via crossbreeding) into the genome of anatomically modern humans, with the first two introgressions resulting from modern humans mating with Neanderthals and Denisovans. The third extinct species is likely a hybrid or close relative of Neanderthals and Denisovans that crossbred with “Out of Africa” modern humans in Asia.
Out of Africa
Around 80,000 years ago, a group of anatomically modern humans left the African continent and migrated to other areas of the world. “We know that from that time onwards, modern humans crossbred with Neanderthals in all the continents, except Africa, and with the Denisovans in Oceania and probably in South-East Asia,” said Jaume Bertranpetit, principal investigator at the Institute of Evolutionary Biology.
According to the Out of Africa model, which is the most accepted demographic scenario for explaining recent human evolution, anatomically modern humans evolved in Africa between 200,000 and 300,000 years ago, then migrated to the rest of the world starting around 50,000 to 80,000 years ago. Recent fossil evidence supports the idea that Homo sapiens left in several waves of migration, as opposed to a sole migratory event.
Once out of Africa, modern humans came upon groups of Neanderthals and Denisovans, with whom they mated. And while we speak of them as species that became extinct, some experts assert that it is more correct to say that they were “absorbed” into our own species through generations of genetic mixing. To this day, our genomes carry the marks of their ancient interactions, including inherited genetic risks and benefits.
Tibetans, for example, have Denisovan gene variants associated with hemoglobin concentration and an adaptive response to hypoxia (lack of oxygen). These gene variants confer advantages to life in high altitudes. “Tibetans utilise five gene variants that collectively make them well adapted to the low oxygen levels, extreme cold, elevated levels of ultraviolet light and limited food supplies that characterise high-altitude living,” wrote Andrew Masterson for Cosmos Magazine.
And celiac disease, a hereditary illness affecting more than one in 100 people, has been linked to known celiac alleles inherited from our Neanderthal ancestors. While celiac disease presents many everyday hurdles for those diagnosed, and is associated with increased chance of death in celiacs who go undiagnosed or misdiagnosed, celiac alleles are also associated with survival advantages. These advantages include increased protection against bacterial infections and dental decay.
Why was deep learning used in the Spanish-Estonian study?
Deep learning methods are useful in scenarios where the data is highly complex and extensive, though the Spanish-Estonian human ancestor study was the first known instance where deep learning was applied to this type of human evolution enquiry.
Deep learning is a subfield of machine learning, and is a specific approach for constructing an artificial neural network that operates in a nonlinear fashion. “The artificial neural networks are built like the human brain, with neuron nodes connected together like a web,” wrote Jake Frankenfield for Investopedia.
The artificial neural network structure allows the computer to analyze data in a similar way that our brains do — taking in many layers of complex information from different sources and storing or “remembering” them in an integrated yet flexible way. This integrated information then acts as a kind of foundation through which new information is interpreted. Deep learning is already being used in language recognition, text generation, chatbots, and personal news aggregation on our phones, to name just a few everyday applications.
The researchers’ unique use of deep learning to discover the third archaic species that has contributed to the modern human genome will pave the way for similar complex enquiries regarding biology and evolution.
Kristen Hovet covers genetics, medical innovations, and the intersection of sociology and culture. Follow her on her website or Twitter @kristenhovet