As weeks became months and the symptom spectrum widened and worsened, researchers began to focus on “host risk factors” to explain the increasingly apparent variability in the COVID-19 experience. According to Jack Kosmicki, PhD, of Regeneron Genetics Center, at the recent American Society of Human Genetics virtual annual meeting:
Genetics is one avenue to better understand why outcomes of COVID are so different. Some patients have so few symptoms that they don’t realize they’re infected, yet the other end of the extreme is requiring hospitalization, or death. Genetic risk factors might influence the likelihood of becoming infected or requiring hospitalization.
So far, very few genes have been linked to COVID-19. Other factors like socioeconomic status, exposure to the virus in the workplace or in crowded housing conditions, being of Black or Asian ancestry and non-genetic pre-existing conditions are more important.
But genetic differences may provide more subtle information, such as accounting for the wildly different and nuanced ways that the new infectious disease unfolds. Perhaps genetic distinctions may explain why some people recover quickly and others become long-haulers, or why some succumb to overwhelming immune responses and others effectively fight off the virus without overdoing it.
Several presentations at the genetics conference converged on a few genome regions that harbor gene variants that affect susceptibility to infection and severity of the illness. They echo reports going back months.
Teasing out genetic associations with a particular COVID outcome requires comparing several types of information about many people. Bioinformaticians seek the needles in the haystacks of human genomes, the sequences of A, T, C, and G that people with similar courses of illness and outcomes uniquely share.
Presenters described a variety of “big data” resources, including consumer DNA testing companies, government biobanks and biopharmaceutical companies. These storehouses provide family trees, health histories, and demographic information. They also include DNA data from cells in cheek scrapings, spit, and blood. The samples hold clues to why one person becomes infected and never knows it, while another progresses rapidly through a horror show of symptoms and complications. Hospital and electronic health records propel many studies too.
Zeroing in on COVID-influencing genes
Determining the DNA sequence of an entire genome, or just the protein-encoding part (the exome), is one way to match genetic information to infectious disease characteristics. But it may also be overkill. Instead many studies use a shortcut called a “GWAS,” for genome-wide association study. A GWAS – less clumsily called an association study – is a little like skimming a book rather than reading every word. The gist is still clear.
An association study catalogs sites in a genome where the DNA base varies in a population. At such a site, called a “single nucleotide polymorphism” or SNP, 97% of a population might have an “A” DNA base, but 2% have a “G” and 1% have a “T,” for example.
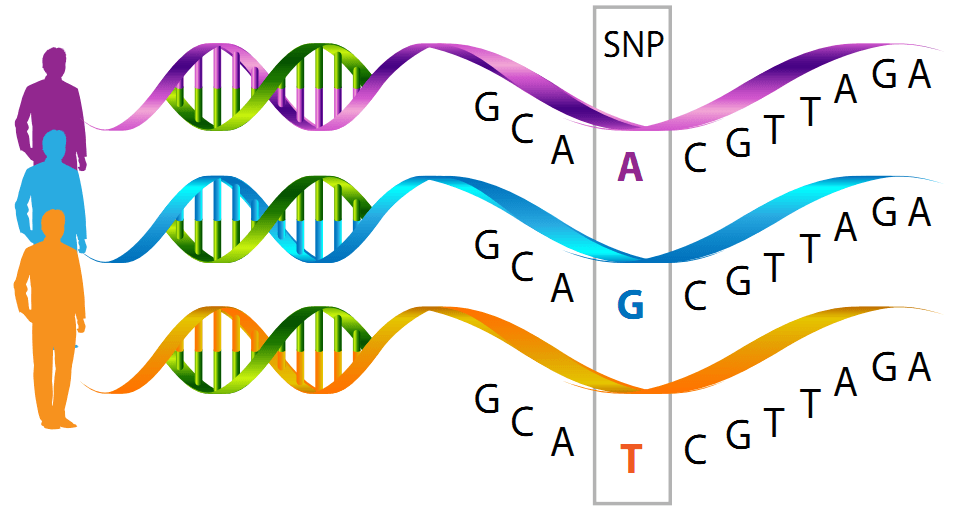
The power of an association study comes from considering thousands of SNPs along a chromosome, which tend to be inherited together, like people in a boat used for crew. Linked SNPs form signatures that distinguish people. The technique is based on the classic genetic concept of ”linkage” of genes on a chromosome.
Here’s the logic. A SNP pattern that is significantly more common among people who share a characteristic, like COVID-19 severe enough to require a ventilator, points to part of a chromosome that might harbor a gene that influences the characteristic. It’s like narrowing down the location of a fleeing burglar to a specific alleyway.
A groundbreaking COVID paper published in June compared SNP patterns among 1,980 patients hospitalized with COVID pneumonia in Italy and Spain during the first wave of the pandemic. The investigators collected data from intensive care units and general wards at seven hospitals in four cities that were pandemic epicenters.
The study tracked 8.5 million SNPs, zeroing in on a set of 13 representing a “gene cluster” in part of chromosome 3. The SNPs are overrepresented among COVID-19 patients who needed hospitalization.
Several other investigations have since replicated the results implicating the chromosome 3 locale. One study that made headlines identified the genetic signature among Neanderthals, which may partly explain why COVID-19 is less prevalent in Africa. The severe-COVID gene variants among Europeans must have arisen, by mutation, after the ancestors of the Neanderthals left Africa 40,000 to 60,000 years ago.
Genetic clues in spit samples and cheek scrapings sent to companies
Consumer DNA testing giants 23andMe and Ancestry DNA conducted association studies in search of COVID commonalities. They tracked infection, respiratory symptoms, hospitalization, demographic factors (age, sex, ethnicity, socioeconomic status) and pre-existing conditions. I gave both companies permission to use my data.
In April and May, AncestryDNA used the data of half a million customers in an association study probing COVID-19 severity. Which patterns of gene variants were more common among the 2,407 people who’d tested positive, or among the 250 who required hospitalization?
The already-identified chromosome 3 gene cluster popped up, plus a region of chromosome 9 near the gene that determines ABO blood type. A possible protective effect of having type O blood had been found in some investigations, but not in others.
In addition, the AncestryDNA researchers found a trio of genetic associations not seen before, in genes that are involved in viral replication (SRRM1), encode an antibody part (immunoglobulin lambda) and control replication of influenza virus (IVNS1ABP).
The variant of IVNS1ABP is particularly interesting because it is associated with COVID-19 susceptibility, but only in males. That might be one reason why evidence is growing that males are more severely affected. But how this happens isn’t known.
The AncestryDNA work provided “new evidence that host genetic variation likely contributes to COVID-19 outcomes and demonstrates the value of large- scale, self-reported data as a mechanism that rapidly address a health crisis,” concluded staff scientist Genevieve Roberts, PhD in a poster presentation.
23andMe’s study also led to the chromosome 3 region and ABO blood type. Slightly more than a million customers signed on for the study, which spanned April 6 through July 25. Of the participants, 15,434 reported a positive COVID-19 test and 1,131 of them were hospitalized.
Perhaps 23andMe’s most compelling finding was that the genetic associations don’t explain the observation that African Americans have a greater risk of hospitalization for COVID-19, after adjusting for education, income, age, sex, obesity, and pre-existing conditions. The results suggest that if there’s a strong genetic explanation for why infection prevalence among people of African ancestry is so much higher than among other population groups, we don’t yet know what it is.
Jack Kosmicki, from Regeneron, and colleagues, reported findings that went beyond an association study, determining the exome sequences of nearly 900,000 individuals. Their data came from the UK Biobank and other resources.
Of the 900,000 people, 13,000 tested positive for COVID. The study also found the chromosome 3 hotspot, and linked the ABO gene to susceptibility but not to severity. The team identified a few other genes, including one that encodes a receptor for type 1 interferon, confirming other studies connecting mutation in this gene to severe COVID-19.
Some investigations reach high numbers by combining other studies, an approach called a meta-analysis. A data-pooling resource for the scientific community, the COVID-19 Host Genetics Initiative, enabled Andrea Ganna, PhD, of the Institute for Molecular Medicine at the University of Helsinki and Kenneth Baillie, PhD, of the University of Edinburgh, Roslin Institute, and their team to evaluate almost 31,000 cases among 1.7 million people from 36 studies done in 16 countries.
Their findings, published in Science and reported at the meeting, also associated types A, B, and AB blood with greater susceptibility to infection, but not necessarily with greater severity. The work revealed a variant in a gene on chromosome 19 associated with interstitial lung disease that may raise risk of developing pneumonia, and another on chromosome 21 associated with the inflammatory response, which can spiral out of control in COVID.
Putting genetic risk into perspective
Many of the abstracts for the recent human genetics conference began with acknowledging the weakness of the evidence for genetic risk factors in COVID-19 susceptibility and severity. Given that the investigations are based on such huge samples, the overall picture points to a larger role for environmental influences than genetic ones, at least in the studied populations, mostly in North America and Europe. There were almost no data from Africa, where some research suggests genetics might play a larger role (see Despite poor healthcare, Africa leads the world in controlling COVID-19. Here are some reasons why). And that’s been what we’ve seen from the beginning.
Let’s look back.
Perhaps the most alarming point in the pandemic was when epidemiologists, public health experts, and physicians and nurses realized that community spread, in King County and Snohomish County, Washington state, stemmed from people who were infected but didn’t have symptoms.
“A series of unfortunate events” reads the abstract for the paper in Science tracing these cases, identified between February 20 and 15 March 15, 2020. The CDC reported the details on March 27 here.
Healthcare workers, particularly those who’d treated patients from the Life Care Center, a skilled nursing facility in Kirkland, tried to warn the rest of the country to pay attention. But the new infectious disease was already spreading throughout the country.
Since then, many studies have documented transmission from asymptomatic people. Estimates of the percentage of infected individuals who don’t have or notice symptoms range from 20% to up to 90%, depending on who’s asked.
One explanation for asymptomatic spread, with building evidence, is that some people have protection from T cells against a past infection with a related coronavirus that causes the common cold. When they encounter SARS-CoV-2, those T cells rapidly rev up an antibody response before symptoms even begin – but the person can still spread the disease.
Another environmental driver of COVID-19 is degree and duration of exposure. A healthy, fit female health care worker with type O blood might have demographics and the meager genetic evidence in her favor, but she isn’t immune from sustained exposure to patients with high viral loads, even with PPE. More than 1,700 healthcare workers have died from the infection.
The emerging picture of a greater contribution of nurture over nature in COVID-19 susceptibility and severity is good news, because we can control some environmental factors, at least to some extent. Until vaccines become widely available, it’s best to continue to follow public health measures to directly limit transmission of the virus.