Imagine traveling to a parallel universe and several centuries into the future. Now let’s say you end up at a golf course to play a round. You don’t have any equipment, so you head to the shop to rent some clubs, and the clerk hands you a bag of baseball bats! You’re thinking, “WTH,” but nobody else bats an eyelash at this absurdity. This is just how the humans around here play golf!
A few years from now, we’ll look back on how we practiced some aspects of medicine and be just as appalled. Human nature is to make the best of the resources we have while perpetually refining our techniques.
Enough with geeky analogies. What is precision medicine?
The topic of precision medicine inevitably comes up when I discuss my job these days. Friends and family usually say something to the effect of, “So, precision medicine makes healthcare more…precise?” Well, yes! Personalized or precision medicine, terms often but not always used interchangeably, are essentially terms for a revolutionary and new era in medicine. The future of healthcare will involve using genomic and other omic information to streamline diagnosis, drug development, prevention, and treatment of human disease and disorders. We don’t precisely know what the future holds, and physicians are still learning the implications, while scientists are still honing “bench to bedside” approaches.
Digressing for a sec. Need to give a primer on genomics and NGS.
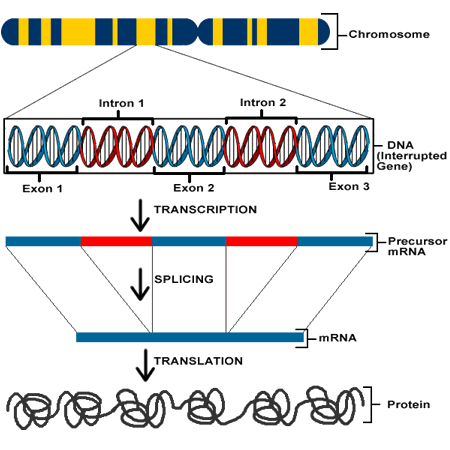
Stealing rather unapologetically from my previous post, here is a briefing for a layperson on genetic literacy: Essentially, proteins are the most basic functional components of living things. Proteins serve all purposes from structure, immunity, metabolic, nutritive, enzymatic functions, and more. They are macromolecules comprised of amino acid chains (polypeptides.) The sequence of amino acids in any protein determines its 3D structure. This sequence of amino acids is determined by codons, each codon coded for by 3 adjacent nucleotides. The DNA in a gene of any organism can be transcribed (into RNA), and translated (into proteins) in many varied permutations by alternative splicing of introns, allowing the functions of life to be carried out. This is a very abridged explanation, but there are some nice primers here and here.
During the Human Genome Project, started in 1990, it took about 10 years and $2-3 billion to sequence a single human genome. Now the newest NGS (Next Generation Sequencing) machines can do the same for a few thousand dollars and in about a day, depending on coverage and other specifics. Basically, we can take a human tissue sample, or tissue samples from large patient cohorts, and get data in the form of a very long sequence comprised of the four nucleotides, A,C, G, and T.
Bench to bedside is used to describe the process of taking these samples from the “lab bench,” and transforming the information into clinically actionable medical treatment at patients’ bedsides.
Because I am only now realizing I could write an entire series on this, I’ll briefly discuss the most exciting aspects of personalized medicine.
GWA Studies:
A GWAS (Genome Wide Association Study) is a relatively new approach to finding genetic markers correlated with disease. A genome is comprised by the entirety of DNA contained within the chromosomes of an organism. Researchers examine 500,000 to a million SNPs across a large sample size of individuals with a given disease compared to a control group to find markers more common in the disease group than the control. SNPs (pronounced “snips”) are individual locations across a genome where nucleotide variations can occur (more than 99 percent of all humans’ genomes are the same!) The human genome is comprised of about 3 billion base pairs, of which only 10 million specific nucleotide loci can vary across the entire human species. Because of linkage disequilibrium, only a handful of these need examining in a GWAS to glean meaningful information. Importantly, SNPs found to be significant in a GWAS are not necessarily the causative variation of a disease; rather they are strongly correlated with the disease sample. GWAS have been the prevalent tool used over the last few years by researchers of complex diseases. Because complex or multi-factorial diseases, like heart disease, diabetes, and many cancers, are influenced by several factors including environment, lifestyle, and genetics, they are harder to understand than the more rarely occurring but predictably inherited Mendelian or single-gene disorders, like Cystic Fibrosis and Sickle-cell anemia.
Diagnostics:
The current and future of diagnostics is so vast, I can’t possibly purport to cover this here. From relatively simple tests for single-gene disorders, to the NIH Undiagnosed Diseases Program, we’re just scratching the surface.
To touch on one example, if you’ve ever been an expecting parent, you may have taken the quad screen, or even a test like Maternit21 to determine whether your unborn fetus suffers from any of a few trisomies that causes diseases like Down syndrome. In the very near future, the whole fetal genome could be sequenced early in pregnancy, bringing a world of information (and a world of ethical questions.)
Another example of the future of diagnostics involves the case of a child with a mysterious gastrointestinal illness that would kill him if doctors couldn’t find a sound treatment. Long story short, the quest was like finding a specific needle in a stack of needles. Through Whole Exome Sequencing, doctors and scientists acted as detectives to solve the case. I encourage everyone to read this amazing Pulitzer Prize-winning story.
Pharmacogenomics:
Pharmacogenomics uses a patient’s genomic information to streamline a drug’s effectiveness while minimizing any adverse reactions.
We’re all familiar with the list of potential side effects listed on pharmaceuticals. For instance with cancer treatment and oncology drugs, chemo has often been a trial and error journey. Today, and more so in the future, patient sequence data will provide powerful information on how s/he will metabolize a drug. While one patient with a disease may have the proper enzyme activity for a drug to be efficacious, another may not respond sufficiently, and thus require a higher dose. In addition, while one patient with a specific disease may metabolize a drug normally, another may metabolize it too slowly or quickly, causing toxic buildup. Yet other patients with certain genetic variations could have adverse reactions to a drug, and shouldn’t take it at all (remember Vioxx?). Pharmacogenomic tests give clinicians this vital knowledge to help make informed treatment decisions. There are already actual apps like iWarfarin in which a physician can plug in patient data including gender, weight, age, and presence or absence of certain biomarkers, and the app spits out an appropriate dose. In the upcoming years, drugs that were previously abandoned during development due to potential adverse reactions may again be researched and released. For the first time in history, drugs may soon require genetic testing prior to prescription.
Okay, now I’m rambling, and I commend you if you’ve gotten this far. There’ll be more to come from me, and far more to come from precision medicine!
NOTE: This article originally appeared in SkepChick–http://skepchick.org/2014/06/guest-post-what-is-precision-medicine-2/–and was provided to GLP by the author and with permission of the original site.