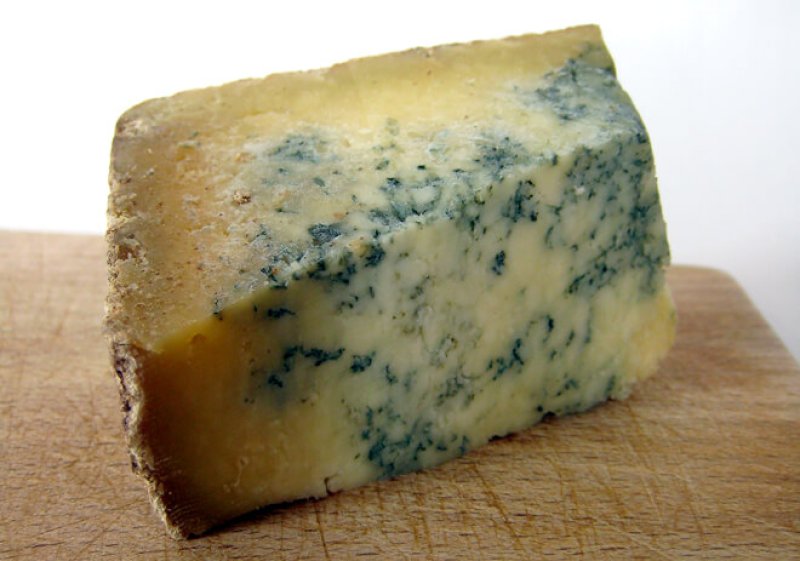
You and your favorite cheese—whether it’s cheddar, Wensleydale, or a good aged goat brie—have something in common: You’re both home to a constantly evolving menagerie of microbes. The bacteria inside you and your fermented dairy live together in a community called a biome, growing and changing in response to their environments. And they adapt to their homes—a cow’s hide, a chunk of Swiss, or your gut—by stealing their neighbors’ genes.
That genetic transfer has the ability to dramatically change a microbe. … In humans, that’s how antibiotic resistance can emerge—one bug evolves a mutation that helps it survive the onslaught of a drug, and it makes its way into the rest of the community. But to fully understand how resistance evolves, studying superbugs isn’t enough: You need large, diverse bacterial boroughs to understand how bugs siphon off new genes.
The search for lively bacterial communities led Rachel Dutton, a microbiologist at UC San Diego, to cave-aged cheese wheels….
That kind of information could help researchers figure out which genes are most prone to transferring within the human microbiome.
The GLP aggregated and excerpted this blog/article to reflect the diversity of news, opinion, and analysis. Read full, original post: What gene-swapping cheese microbes could say about antibiotic resistance