The emerging field of cyberbiosecurity explores the whole new category of risks that come with the increased use of computers in the life sciences.
University scientists, industry stakeholders and government agents have begun gathering to discuss these threats. We’ve even hosted FBI agents from the Weapons of Mass Destruction Directorate here at Colorado State University and previously at Virginia Tech for crash courses on synthetic biology and the associated cyberbiosecurity risks. A year ago, we participated in a U.S. Department of Defense-funded project to assess the security of biotechnology infrastructures. The results are classified, but we disclose some of the lessons learned in our new Trends in Biotechnology paper.
Along with co-authors from Virginia Tech and the University of Nebraska-Lincoln, we discuss two major kinds of threats: sabotaging the machines biologists rely on and creating dangerous biological materials.
Computer viruses affecting the physical world
In 2010, a nuclear plant in Iran experienced mysterious equipment failures. Months later, a security firm was called in to troubleshoot an apparently unrelated problem. They found a malicious computer virus. The virus, called Stuxnet, was telling the equipment to vibrate. The malfunction shut down a third of the plant’s equipment, stunting development of the Iranian nuclear program.
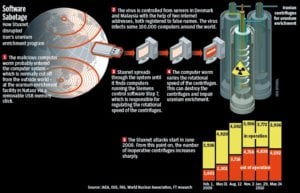 Unlike most viruses, Stuxnet didn’t target only computers. It attacked equipment controlled by computers.
Unlike most viruses, Stuxnet didn’t target only computers. It attacked equipment controlled by computers.
The marriage of computer science and biology has opened the door for amazing discoveries. With the help of computers, we’re decoding the human genome, creating organisms with new capabilities, automating drug development and revolutionizing food safety.
Stuxnet demonstrated that cybersecurity breaches can cause physical damages. What if those damages had biological consequences? Could bioterrorists target government laboratories studying infectious diseases? What about pharmaceutical companies producing lifesaving drugs? As life scientists become more reliant on digital workflows, the chances are likely rising.
Messing with DNA
The ease of accessing genetic information online has democratized science, enabling amateur scientists in community laboratories to tackle challenges like developing affordable insulin.
But the line between physical DNA sequences and their digital representation is becoming increasingly blurry. Digital information, including malware, can now be stored and transmitted via DNA. The J. Craig Venter Institute even created an entire synthetic genome watermarked with encoded links and hidden messages.
Twenty years ago, genetic engineers could only create new DNA molecules by stitching together natural DNA molecules. Today scientists can use chemical processes to produce synthetic DNA.
The sequence of these molecules is often generated using software. In the same way that electrical engineers use software to design computer chips and computer engineers use software to write computer programs, genetic engineers use software to design genes.
That means that access to specific physical samples is no longer necessary to create new biological samples. To say that all you need to create a dangerous human pathogen is internet access would be an overstatement – but only a slight one. For instance, in 2006, a journalist used publicly available data to order a fragment of smallpox DNA in the mail. The year before, the Centers for Disease Control used published DNA sequences as a blueprint to reconstruct the virus responsible for the Spanish flu, one of the deadliest pandemics of all time.
With the help of computers, editing and writing DNA sequences is almost as easy as manipulating text documents. And it can be done with malicious intent.

First: Recognize the threat
The conversations around cyberbiosecurity so far have largely focused on doomsday scenarios. The threats are bidirectional.
On the one hand, computer viruses like Stuxnet could be used to hack into digitally controlled machinery in biology labs. DNA could even be used to deliver the attack by encoding malware that is unlocked when the DNA sequences are translated into digital files by a sequencing computer.
On the other hand, bad actors could use software and digital databases to design or reconstruct pathogens. If nefarious agents hacked into sequence databases or digitally designed novel DNA molecules with the intent to cause harm, the results could be catastrophic.
And not all cyberbiosecurity threats are premeditated or criminal. Unintentional errors that occur while translating between a physical DNA molecule and its digital reference are common. These errors might not compromise national security, but they could cause costly delays or product recalls.
Despite these risks, it is not unusual for researchers to order samples from a collaborator or a company and never bother to confirm that the physical sample they receive matches the digital sequence they were expecting.
Infrastructure changes and new technologies could help increase the security of life science workflows. For instance, voluntary screening guidelines are already in place to help DNA synthesis companies screen orders for known pathogens. Universities could institute similar mandatory guidelines for any outgoing DNA synthesis orders.
There is also currently no simple, affordable way to confirm DNA samples by whole genome sequencing. Simplified protocols and user-friendly software could be developed, so that screening by sequencing becomes routine.
The ability to manipulate DNA was once the privilege of the select few and very limited in scope and application. Today, life scientists rely on a global supply chain and a network of computers that manipulate DNA in unprecedented ways. The time to start thinking about the security of the digital/DNA interface is now, not after a new Stuxnet-like cyberbiosecurity breach.
Jenna Gallegos is a Posdoctoral Researcher in Chemical and Biological Engineering at Colarado State University. Jenna got her PhD in plant biotechnology at UC Davis where she studied gene expression in plants.
Jean Peccoud is a Professor in Synthetic Biology at Colorado State University. Dr. Peccoud’s research program focuses on synthetic biology informatics. His group combines computational and experimental efforts to develop predictive models of behaviors encoded in synthetic DNA sequences. He is the founding Editor-in-Chief of the journal Synthetic Biology published by Oxford University Press.
This article was originally published on the Conversation’s website as “DNA has gone digital – what could possibly go wrong?” and has been republished here with permission.