A tale of a teen and a tyke with the extremely rare Wiedemann Steiner Syndrome (WSS) argues for the value of earlier exome sequencing in the search for a diagnosis.
 Monica’s story: 14 years to diagnosis
Monica’s story: 14 years to diagnosis
Monica was born at the end of 1999. Her small size, early teeth, wide-set eyes, low-set ears, sleep apnea, small mouth with large tonsils and adenoids that blocked her airway, small rotated kidney, and fused vertebrae and spinal disc problems suggested a genetic syndrome. But because she met developmental milestones, genetic testing was minimal: a chromosome check during her third year, and then a microarray test for short pieces of missing or extra chromosomes. All normal.
But as Monica grew, she experienced speech delay, mild developmental delay and intellectual disability, learning disabilities and continuing breathing difficulties. Speech therapy, math tutors and a social worker to help her manage anxiety in social situations enabled her to explore interests, “finally settling into dance and gymnastics for fun and fitness,” her mother Tracey Carmichael recalled. But something wasn’t quite right.
By the time Monica was eight, the pediatrician diagnosed Opitz G/BBB syndrome, based on symptoms that would turn out to be a subset of WSS: mild intellectual disability and developmental delay, speech delay and the wide eyes and low ears. An autism diagnosis didn’t come until 2017, but it was likely present earlier, and part of both syndromes.
“The Opitz diagnosis allowed Monica to access additional funded services at school and tax benefits to offset extra costs at home for tutoring and therapies. It served her well until puberty, when new medical problems came flying at us,” said Tracey, who lives with her daughter in Canada. The new challenges included dysautonomia (autonomic nervous system symptoms), painful joint partial dislocations, poor stomach motility and depression. “It became apparent that her health concerns were not consistent with the Opitz diagnosis, and we asked to be referred back to the geneticist.”
 The diagnosis was wrong, and Monica found the turnaround distressing:
The diagnosis was wrong, and Monica found the turnaround distressing:
When the doctors told me it wasn’t Opitz G/BBB I felt sad and lost because it meant more investigation and more tests, more doctors’ appointments, and time missing school. I was angry at the doctors talking to my parents in terms I didn’t understand. When I asked what they were talking about, the doctors spoke to my parents and not me.
They were back to square one. “I believe that having no diagnosis is infinitely worse than having the wrong diagnosis. A genetic diagnosis should lead to better outcomes through better understanding of the symptoms and better treatment,” Tracey said.
Fortunately, the reanalysis lead to Clara Van Karnebeek at the University of British Columbia, the TIDE BC program seeking genetic causes of intellectual disability and developmental delay, and the correct diagnosis. Participating in research was the only way to have exome sequencing in 2014, and Monica had it in September of that year. “Wiedemann-Steiner Syndrome surprised everyone,” Tracey said.
Today Monica accepts her diagnosis: “It’s a unique part of who I am. I’m trying to take better care of myself by going to my doctors, taking medications, going to rehab and physiotherapy every week and doing exercise at home every day, doing the things I have to do to feel better.”
The diagnosis: WSS
A flurry of case reports of what would be named Wiedemann-Steiner Syndrome appeared in the medical literature in 1989, the name honoring two of the researchers. Affected individuals had long lashes, wide-set eyes, arched brows, a long philtrum (the space between the nose and the upper lip), short nose, low-set ears and a high palate. They had delayed speech, hypotonia (floppiness), and some had intellectual disability and/or developmental delay. The young patients were all short and stocky. Less common were hairy elbows, whorls of hair on the upper back, a dimple in the lower back and spaces in the kidneys.
More reports trickled in, the chromosomes always normal. Then in 2012 Wendy Dawn Jones, at the University of London, showed that WSS typically arises as a new mutation, after which it’s passed on in an autosomal dominant manner; each child of an affected person would have a 50-50 risk. Her team sequenced the exomes of four patients, three of whom had mutations in the same gene, KMT2A. The fourth had slightly different symptoms and facial features, and a mutation in a different gene.
The mutations that cause WSS shorten an enzyme (lysine methyltransferase 2A) that affects the pattern in which two small organic (carbon-containing) groups – methyl and acetyl – attach to certain types of histone proteins around which DNA wraps. Methyls, acetyls, and phosphates control which genes are exposed enough to be expressed, and which aren’t, the chemical crux of epigenetics. So a mutation that affects a crucial epigenetic enzyme alters the expression of several genes. Mutations in other histone-methylation genes produce similar symptoms, such as short stature, speech delay, and distinctive faces, including Kabuki, Rubinstein-Taybi, and floating harbor syndromes.
In 2016 a different research group added mutations behind WSS that substitute an amino acid in the enzyme rather than stunting it, and these individuals also have seizures and immunodeficiency. In 2017 Dr. Jones’s PhD thesis (she was already an MD) added 84 more cases.
An interesting aside: KMT2A has another name. It’s also known as the MLL gene that lies behind mixed lineage leukemia. In WSS the gene is shortened in all cells; in MLL it is cut out and moved to another chromosomal location, landing in a cancer-causing gene, but only in white blood cells.
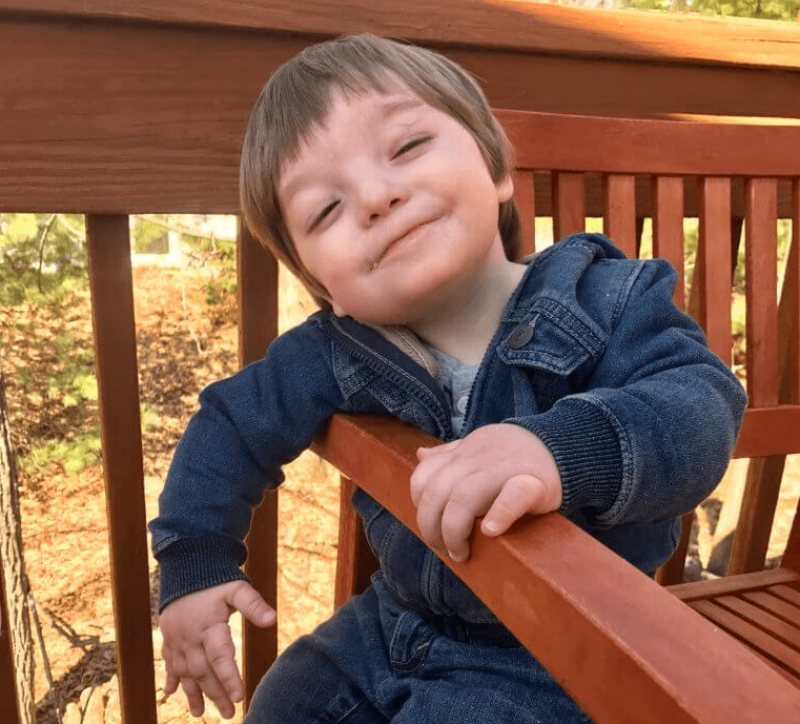
Nicholas’s story: 2 years to diagnosis
By the time Nicholas Lagravinese was born in August 2015, genetic diagnosis had evolved from chromosome checks and microarrays to exome sequencing. But the new technique was still part of research protocols. The little boy seemed fine at birth, but once home he didn’t gain weight, and then began to lose weight – dramatically. In his first weeks, medical attention focused on heart and respiratory problems, and he had surgery to close a hole in his heart.
 At Morgan Stanley Children’s Hospital in Manhattan, the boy was seen by a parade of specialists, including a geneticist who recognized a possible genetic syndrome in the unusual facial features, heart anomalies, aspiration issues and the old-fashioned but still-telling “failure to thrive.” He suggested that Nicholas and his parents have chromosomal microarray testing and exome sequencing. Nicholas was only two months old when the trio gave blood for these tests. If the condition was recessive, the parents would be carriers, of something. But they weren’t.
At Morgan Stanley Children’s Hospital in Manhattan, the boy was seen by a parade of specialists, including a geneticist who recognized a possible genetic syndrome in the unusual facial features, heart anomalies, aspiration issues and the old-fashioned but still-telling “failure to thrive.” He suggested that Nicholas and his parents have chromosomal microarray testing and exome sequencing. Nicholas was only two months old when the trio gave blood for these tests. If the condition was recessive, the parents would be carriers, of something. But they weren’t.
The microarray test came back normal quickly, and the exome results were in by February 2016. Two gene variants stood out as possibly explaining Nicholas’ symptoms, one indicating Wiedemann-Steiner Syndrome. Further analysis as part of a research program nailed the diagnosis by April 2017. Nicholas had a dominant mutation that originated in him.
Remembers Kim Hoffman, Nicholas’s mother, “Receiving the news of the official diagnosis was difficult for us initially. But we are so grateful for the advancements in genetic testing allowing us to learn of this diagnosis so early because it only helps us help Nicholas better; and, Nicholas is still our same lovable little boy that he was before the diagnosis.”
Placing a name on a spectrum of symptoms has eased getting services, such as attending a special needs school and receiving physical, feeding, speech, and occupational therapies, plus a monthly meeting with a nutritionist. Nicholas continues to see the many specialists required to address his associated medical conditions. He’s had five surgeries.
 “Nicholas is developmentally delayed, but from his therapies, school, and a lot of hard work at home, is making great progress. We don’t take any milestones for granted in our house, celebrating all the victories, whether it’s putting pieces in a puzzle, taking a sip out of an open cup, acting out ‘happy and you know it’ and ‘itsy bitsy spider,’ or learning a new sign to communicate. We’ll never forget the big smile plastered on his face and his vocalizing pleasure the first time he walked 50 steps in a row across our living room. We cried tears of joy,” Kim said.
“Nicholas is developmentally delayed, but from his therapies, school, and a lot of hard work at home, is making great progress. We don’t take any milestones for granted in our house, celebrating all the victories, whether it’s putting pieces in a puzzle, taking a sip out of an open cup, acting out ‘happy and you know it’ and ‘itsy bitsy spider,’ or learning a new sign to communicate. We’ll never forget the big smile plastered on his face and his vocalizing pleasure the first time he walked 50 steps in a row across our living room. We cried tears of joy,” Kim said.
The WSS Foundation introduced Kim, her husband Matt, and Nicholas and little sister Emily to other families. “This has been invaluable and truly life-changing. Our WSS family provides an abundance of knowledge and support. We have already learned so much about how to help Nicholas to become the best version of himself, medically, educationally, and socially,” Kim said.
Bigger picture
As time goes on, exome sequencing will yield more single-gene tests, while continuing to solve medical mysteries. Many argue for it to be moved up in the diagnostic process.
A telling investigation came from Tiong Yang Tan, PhD, at the Victorian Clinical Genetics Services in Melbourne, Australia, and colleagues. They diagnosed 23 of 44 kids (up to age 18) from exome sequences, taking only a few months each. The children had undergone many tests and evaluations for on average six years, more than half receiving general anesthesia for a diagnostic procedure. As with Nicholas and Monica, chromosomal checks had turned up nothing, and pediatricians had sent their patients to the exome program because of unusual facial features and multiple anomalies.
The exome sequencing brought surprises. Eight of the 23 kids had mutations in genes that physicians hadn’t even considered. And in six cases the diagnosis led to changed treatment. Furthermore, sequencing the exome before all the standard testing, when signs and symptoms suggest a single-gene condition, would save on average $6,840 per patient, the researchers concluded.
When exome (or genome) sequencing becomes the standard of care for diagnosing mysterious syndromes, the time to diagnosis for even the rarest of genetic diseases will take just hours – like a flight from New York City to London.
(Thanks to the Lagravinese family and Monica for photos, and for the WSS Foundation for contacting me.)
For further information see:
Wiedemann Steiner Syndrome Parents Network
WSS Foundation Facebook Page
Ricki Lewis is the GLP’s senior contributing writer focusing on gene therapy and gene editing. She has a PhD in genetics and is a genetic counselor, science writer and author of The Forever Fix: Gene Therapy and the Boy Who Saved It, the only popular book about gene therapy. BIO. Follow her at her website or Twitter @rickilewis.