Several years ago, Richard Neher, an evolutionary biologist at the University of Basel in Switzerland, and his colleagues wanted to monitor changes to the flu’s genetic makeup to see if the data would help scientists build more-effective flu vaccines. They developed an online interface … and published the results in a publicly available interactive web browser.
…
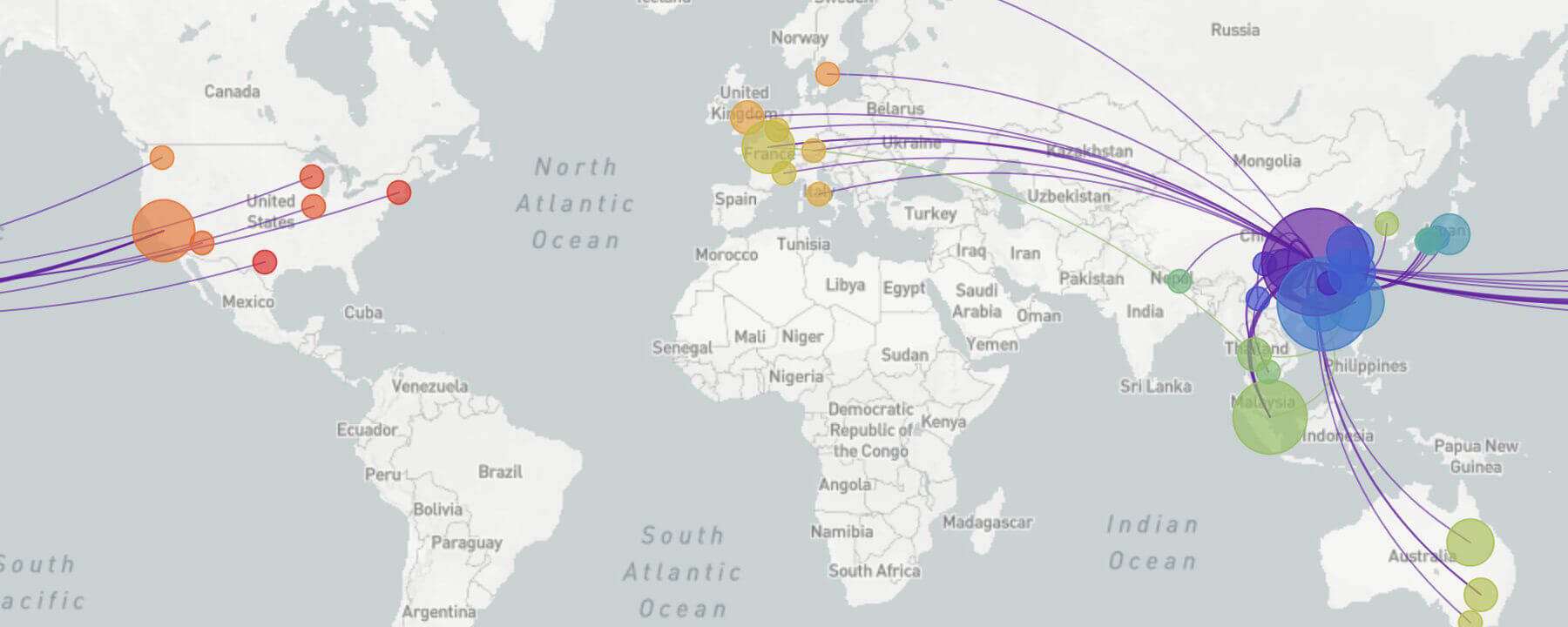
Now, they’ve adapted it to keep track of the genetic tweaks to SARS-CoV-2 as … the virus moves from major hotspots in China to smaller pockets in other countries.
The Scientist: How do viral genomes’ sequences from swabs taken from infected patients help you build a family tree of the virus?
Richard Neher: These coronavirsuses tend to change their genome, they mutate, at a fairly high rate. … As time goes on, the lineages pick up independent mutations, and then they cause outbreaks in different parts of the world.
…
TS: What can the data tell you about the virus’s origins?
RN: The first takeaway is that all these sequences are very, very similar, about eight mutations different than the root. That’s eight mutations in a 30,000-base sequence. What this tells us is that the virus came from one source, not too long ago, somewhere between mid-November and early December.