The controversy over genetically engineered organisms (sometimes called “genetically modified organisms,” or “GMOs”) is genuine, not faux — but only because of uninformed, exaggerated concerns about the most recent techniques’ “newness.” What is faux and disingenuous are the arguments put forth by genetic engineering’s opponents.
Humans have been modifying the DNA of our food for thousands of years (even though we didn’t know that DNA was mediating the changes until the 20th century). We call it agriculture. Early farmers (>10,000 years ago) used selective breeding to guide DNA changes in crops and animals to better suit our needs. Approximately a hundred years ago plant breeders began using harsh chemicals and/or radiation to randomly change, or mutate, the DNA of crop plants. These mutagens caused innumerable changes to the DNA, none of which was characterized or examined for safety. The plant breeders looked and selected for desired traits of various kinds. Problems were rare.
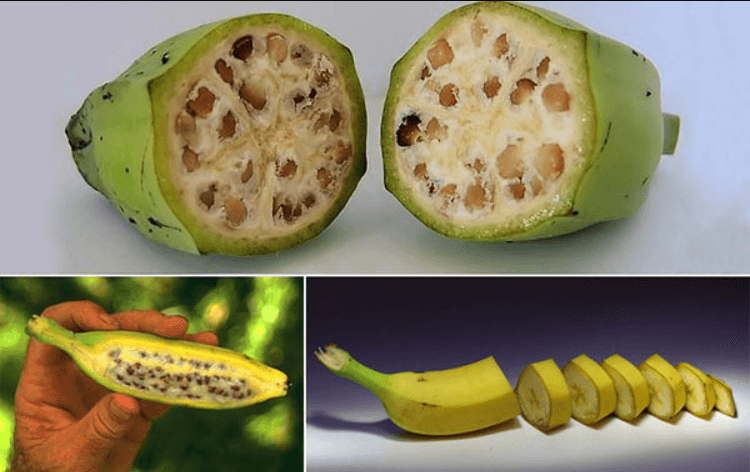
Today more than half of all food crops have mutagenesis breeding as part of their pedigree. Ancestral varieties bear little resemblance to the domesticated crops we eat today. There are many striking pictorial examples here.

Approximately 40 years ago agricultural scientists and plant breeders began to use recombinant DNA technology (“gene splicing”) to make far more precise and predictable changes to the DNA in our crop plants. This molecular genetic engineering (GE) typically takes a gene with a known function, e.g., one that expresses a protein toxic to certain insect predators, and transfers it into a crop, enabling the GE crop to protect itself from insect pests. This one trait, resulting from the introduction of a gene from the bacterium Bacillus thuringiensis (abbreviated “Bt”) into plants, has allowed farmers around the world to reduce broad spectrum insecticide spraying by billions of pounds.
Since the advent in the 1970s of this recombinant DNA technology, which enables segments of DNA to be moved readily and more precisely from one organism to another, molecular genetic engineering techniques have become increasingly more sophisticated, precise, and predictable. This evolution has culminated in the most recent discoveries, the CRISPR-Cas9 system and base editing.
CRISPR (short for Clustered Regularly Interspaced Short Palindromic Repeats) is a natural defense system that bacteria use against invading viruses. CRISPR can recognize specific DNA sequences, while the enzyme Cas9 cuts the DNA at the recognized sequence. As often happens in science–and reminiscent of mutagenesis a century ago and recombinant DNA technology in the 1970s–molecular biologists quickly copied, adapted, and improved the naturally occurring system. Using CRISPR-Cas9, scientists can target and edit DNA at precise locations, deleting, inserting, or modifying genes in microorganisms, plants and animals, and even humans.
CRISPR-Cas9 presages a revolution in agriculture and human medicine because it is so much more precise and predictable than earlier techniques. Precision and predictability are important to ensure that results are safe and achieve their desired ends. There are notable historical examples of older, pre-molecular techniques of genetic modification in agriculture that misfired. Examples include:
- The Lenape potato, which contained elevated, harmful levels of a plant alkaloid
- The creation of hyper-aggressive Africanized honeybees by crossbreeding African and European species in the 1950s
- Inadvertently making some varieties of corn in the United States more susceptible to the Southern Corn Leaf Blight fungus, which resulted in significant losses of the U.S. harvest in 1970.
Despite their success for farmers of all types, from subsistence to huge-scale commercial, GE crops have been discriminated against by regulators and demonized by activists. In the early 1970s, at a conclave now referred to as the Asilomar Conference, a group of scientists — none involved in agriculture or food science — raised concerns about hypothetical hazards that might arise from the use of the newly discovered molecular genetic modification technique — recombinant DNA technology, or “gene-splicing.” However, they failed to appreciate the history of genetic modification by means of cruder, less predictable technologies, described below.
The Asilomar Conference led to guidelines published by the U.S. National Institutes of Health (NIH) for the application of these techniques for any purpose. These “process-based” guidelines, which were applicable exclusively to recombinant DNA technology, were in addition to the preexisting “product-focused” regulatory requirements of other federal agencies that had statutory oversight of food, drugs, certain plants, pesticides, and so on.
The NIH guidelines, which were in effect the “original sin” of precautionary, unscientific regulation, were quite stringent. For example, without regulatory approval, the “intentional release” of recombinant DNA-modified organisms into the environment, or fermentation (in contained fermenters) at volumes greater than ten liters, required explicit prior approval by the NIH and local Institutional Biosafety Committees.
Given the seamless continuum of techniques for genetic modification described above, such requirements were unwarranted. No analogous blanket restrictions existed for similar or even virtually identical plants, microorganisms, or other organisms modified by traditional techniques, such as chemical or irradiation mutagenesis or wide-cross hybridizations.

Thus, uninformed, ill-founded, and exaggerated concerns about the risks of recombinant DNA-modified organisms in medical, agricultural, and environmental applications precipitated the regulation of recombinant organisms—regulation triggered simply by the “process,” or technique, of genetic modification, rather than the “product,” i.e., the characteristics of the modified organism itself. This was an unfortunate precedent–as was entrusting technology regulation to a research agency, the NIH, whose legacy plagues regulation worldwide today. Most industrial countries, including the US, have specific regulatory agencies like the US Food and Drug Administration and European Medicines Agency that regulate product safety. Research agencies rarely are involved in regulating products or processes.
The regulatory burden on the use of recombinant DNA technology—which some people (mostly activists and regulators) consider gives rise to a mythical category of organisms called “Genetically Modified Organisms,” or “GMOs,” was, and remains, disproportionate to its risk, and the opportunity costs of regulatory delays and expenses are formidable. According to Wendelyn Jones at DuPont Crop Protection, a survey found that “the cost of discovery, development and authorization of a new plant biotechnology trait introduced between 2008 and 2012 was $136 million. On average, about 26 percent of those costs ($35.1 million) were incurred as part of the regulatory testing and registration process.”
A salient question currently for regulators, scientists, and consumers is whether gene editing will fall down the same rabbit hole. Unfortunately, much of the discussion focuses on irrelevant issues such as whether organisms that could arise “naturally” or that are not “transgenic” (containing DNA from different sources) should be subject to more lenient regulation than “GMOs.” As should be evident from the discussion above, such issues have no implications for risk—and, therefore, for regulation. In fact, modern plant breeding techniques, including genome editing, are more precise, circumscribed, and predictable than other methods — in other words, if anything, likely to be safer. This assessment is neither new nor novel. A landmark report from the U.S. National Research Council concluded in 1989:
- Information about the process used to produce a genetically modified organism is important in understanding the characteristics of the product. However, the nature of the process is not a useful criterion for determining whether the product requires less or more oversight.
- The same physical and biological laws govern the response of organisms modified by modern molecular and cellular methods and those produced by classical methods.
- Recombinant DNA methodology makes it possible to introduce pieces of DNA, consisting of either single or multiple genes, that can be defined in function and even in nucleotide sequence. With classical techniques of gene transfer, a variable number of genes can be transferred, the number depending on the mechanism of transfer; but predicting the precise number or the traits that have been transferred is difficult, and we cannot always predict the phenotypic expression that will result. With organisms modified by molecular methods, we are in a better, if not perfect, position to predict the phenotypic expression.
- With classical methods of mutagenesis, chemical mutagens such as alkylating agents modify DNA in essentially random ways; it is not possible to direct a mutation to specific genes, much less to specific sites within a gene. Indeed, one common alkylating agent alters a number of different genes simultaneously. These mutations can go unnoticed unless they produce phenotypic changes that make them detectable in their environments. Many mutations go undetected until the organisms are grown under conditions that support expression of the mutation.
- [N]o conceptual distinction exists between genetic modification of plants and microorganisms by classical methods or by molecular techniques that modify DNA and transfer genes.
These critical points, clearly articulated more than 30 years ago and about which there is virtual unanimity in the scientific community, have not sunk in.

There is an ongoing need for genetic modification in agriculture. Gene editing could play a key role in England’s sugar beet sector, for example, and Britain’s farming and in February, environment minister George Eustice told the annual conference of the National Farmers Union that the sugar beet sector could use the assistance of gene editing technologies to overcome yield reduction due to virus infection. He added: “Gene editing is really just a more targeted, faster approach to move traits from one plant to another but within the same species so in that respect it is no different from conventional breeding.”
The first part of Eustace’s statement is accurate, but the second part gives the misimpression that although gene edited crops are analogous to “conventional breeding”—and, therefore, presumably harmless—they are sufficiently far removed from dreaded GMOs that they should be exempt from the onerous regulation appropriate for the latter. Until now, in the European Union, gene editing has been strictly regulated in the same way as GMOs. Their oversight might diverge in the future, however, inasmuch as serious attention is being paid to this new technology and its enormous potential.
But preferential regulatory treatment of gene editing over recombinant DNA-mediated modifications would represent expediency over logic: The NRC report (as well as other, innumerable, similar analyses) makes it clear that an approach that deregulates gene editing but not recombinant DNA modifications would ignore the seamless continuum that exists among methods of genetic modification, and that it would be unscientific. There is no reason to throw transgenic recombinant DNA constructions under the regulatory bus.
The relationships among genome editing, plant breeding, and GMO crops are more interconnected, complex and nuanced than it may appear at first glance. Plant breeding itself has long been a murky science in terms of genetics and heredity. While Britain’s Eustice lauds genome editing because it involves only intra-species modification, the history of plant breeding has long included “distant” or “wide” crosses to move beneficial traits such as disease resistance from one plant species or one genus to another. Almost a century of wide cross hybridizations, which involve the movement of genes from one species or genus to another, has given rise to plants–including everyday varieties of corn, oats, pumpkin, wheat, black currants, tomatoes, and potatoes, among others–that do not and could not exist in nature. Indeed, with the exception of wild berries, wild game, wild mushrooms, and fish and shellfish, virtually everything in North American and European diets has been genetically improved in some way. Compared to the new molecular modification technologies, these wide crosses are crude and less predictable.
Another wrinkle is that plant scientists have discovered what have been termed “natural GMOs,” which further confounds the terminology. These include whiteflies harboring plant genes that protect them from pesticides, horizontal gene transfer between different species of grasses, sweet potato harboring sequences from the bacterium Agrobacterium, and aphids which express a red fungal pigment to protect them from would-be predators. This is more evidence that the term “GMO” itself has become meaningless.
This brings us back to the regulatory conundrum surrounding the way forward with the various products of genetic engineering using different technologies. Eager to avoid the delays, impasses and rejections — and inflated opportunity costs — that have confounded “GMOs,” many in the scientific and commercial communities are willing to play down the novelty of genome editing, while, in effect, conceding that recombinant DNA constructions should continue to be stringently regulated.
However, as we have discussed, the comparison of genome editing and recombinant DNA is a distinction without a difference, especially when viewed against the backdrop of the crude constructions of (largely unregulated) traditional plant breeding. Trying to draw meaningful distinctions between molecular genetic engineering and other techniques for the purpose of regulation is rather like debating how many angels can dance on the head of a pin. It’s way past time that for purposes of regulatory policy, we began to think in terms of the risk posed by organisms and their products, rather than which technology(ies) was employed.
Kathleen Hefferon, Ph.D., teaches microbiology at Cornell University. Find Kathleen on Twitter @KHefferon
Henry Miller, a physician and molecular biologist, is a senior fellow at the Pacific Research Institute. He was a Research Associate at the NIH and the founding director of the U.S. FDA’s Office of Biotechnology. Find Henry on Twitter @henryimiller